

Microsatellites
Also called simple sequence repeats (SSRs), these are tandemly arranged blocks of short nucleotide sequences, usually 1-10 nucleotides long (though more typically 2 or 3), repeated up to 50 times. The number of repeat units in the block can vary noticeably between individuals within a species. This variation can be targeted by PCR, by placing the primers either side of the block. This leads to highly reproducible, co-dominant, easily analyzed and polymorphic markers. As a result, SSRs represent one of the most widely used markers in MAB.
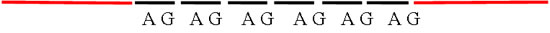 \
\
The di-nucleotide motif “AG” is repeated 6 times in this example of a microsatellite. Primers would be designed from the sequence of the red flanking sequences.
(See Powell et al. 1996)

