Overview of Molecular Breeding Planner
The Molecular Breeding Planner facilitates planning for population development, selections, and crosses using molecular marker data. Marker-assisted selection (MAS) can speed the breeding process by increasing the frequency of favourable alleles and decreasing the frequency of unfavourable alleles in breeding populations. The Molecular Breeding Planner helps breeders develop breeding schemes with timelines to improve the efficiency of genetic gain using genetic markers.
Planning tools support
- Marker-Assisted Recurrent Selection (MARS)
- Marker-Assisted Backcross (MABC)
- Marker-Assisted Gene Pyramiding
Launch
The Molecular Breeding Breeding Planner is a standalone application that can be launched from the Workbench menu.
Alternatively, the Molecular Breeding Planner can be launched independently of the Workbench by accessing the application within the Tools folder.
Interface
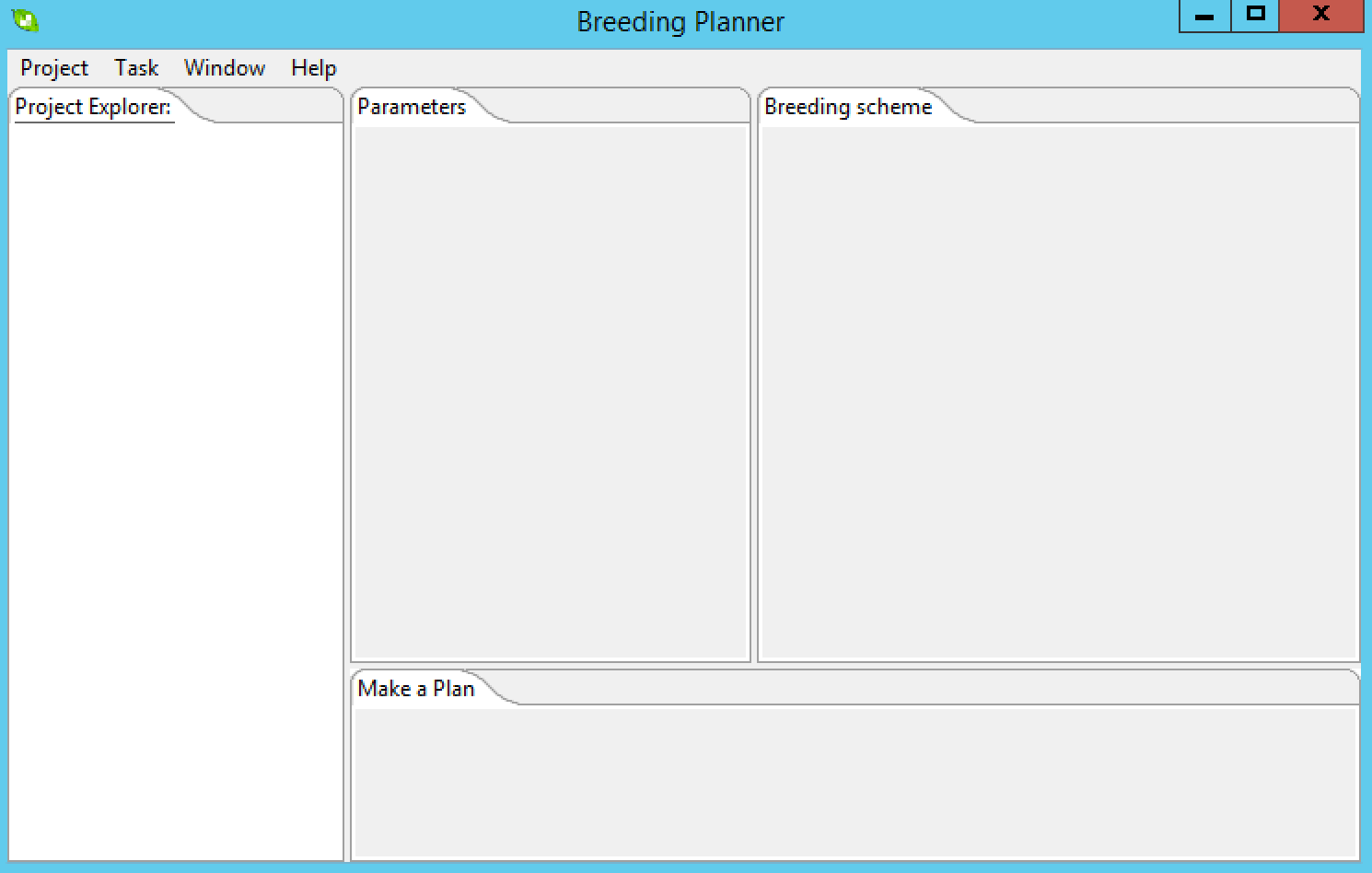
The interface contains the following windows, which will be empty until you establish a project:
- Project Window: Lists all molecular breeding programs within an open project- three distinct programs can be considered: MARS, MABC and MAS for gene pyramiding.
- Parameter Setting/Viewing Window: You can edit/view your breeding parameters in this window.
- Breeding Scheme Window: Once the breeding parameters are specified, a breeding flowchart will be demonstrated in this window.
- Plan-Making Window: You can select the current stage/generation of your breeding programs in this window. A detailed plan for the near future will be made by the Molecular Breeding Planner.
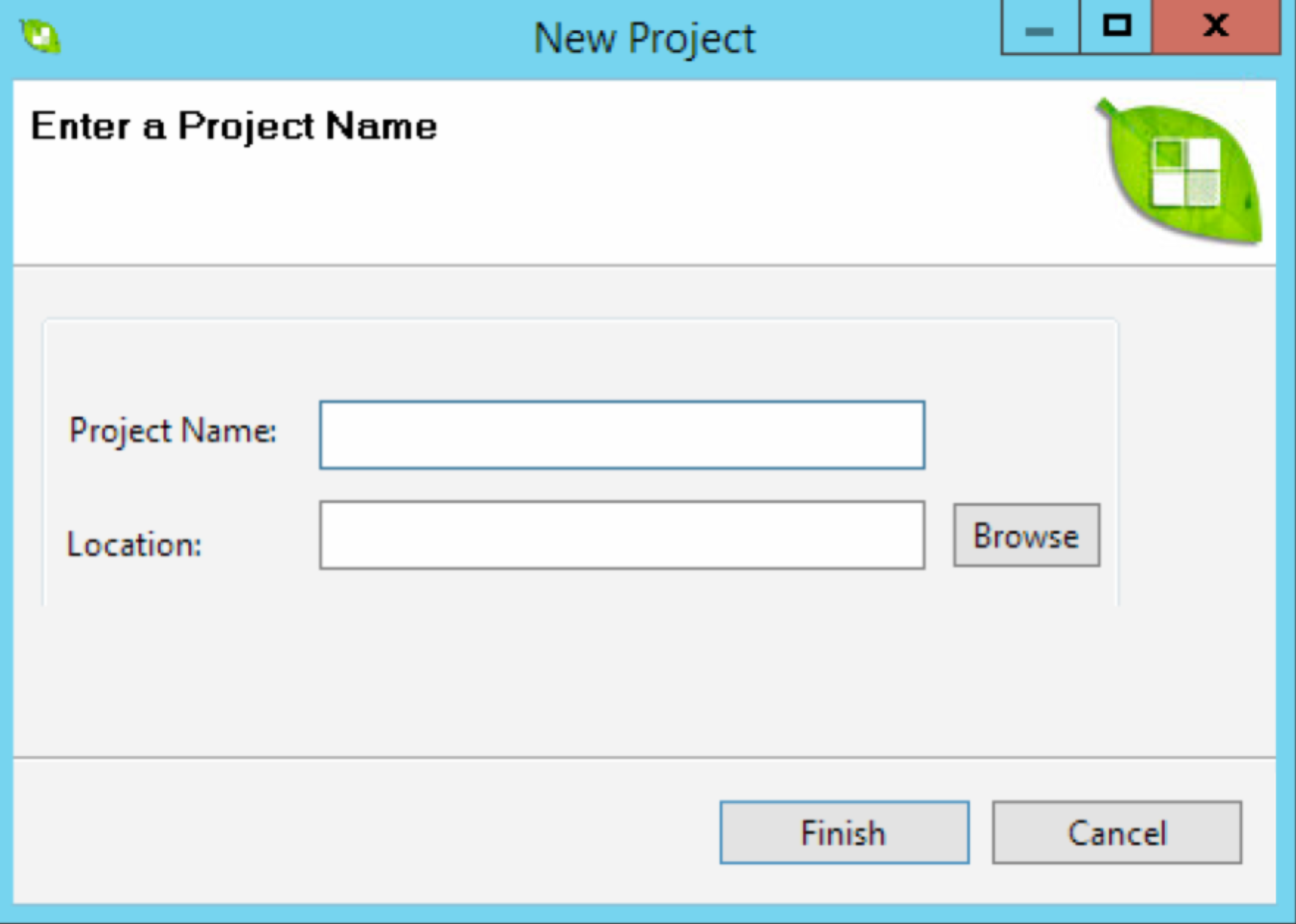
Create a New Project
Select New Project from the Project dropdown menu. Give your project file a name and select a destination folder for the output. Multiple breeding plans can be saved within a single project file.
From the task menu, select one of the new breeding plans:
- MARS: Marker-Assisted Recurrent Selection
- MABC: Marker-Assisted Backcross Breeding
- MAS: Marker-Assisted Selection for Gene Pyramiding
Add details of the breeding plan to the Input Information screen. Users can import the parameters from an external file (file extensions .mars, .mabc, or .mas) or set the parameters by hand.
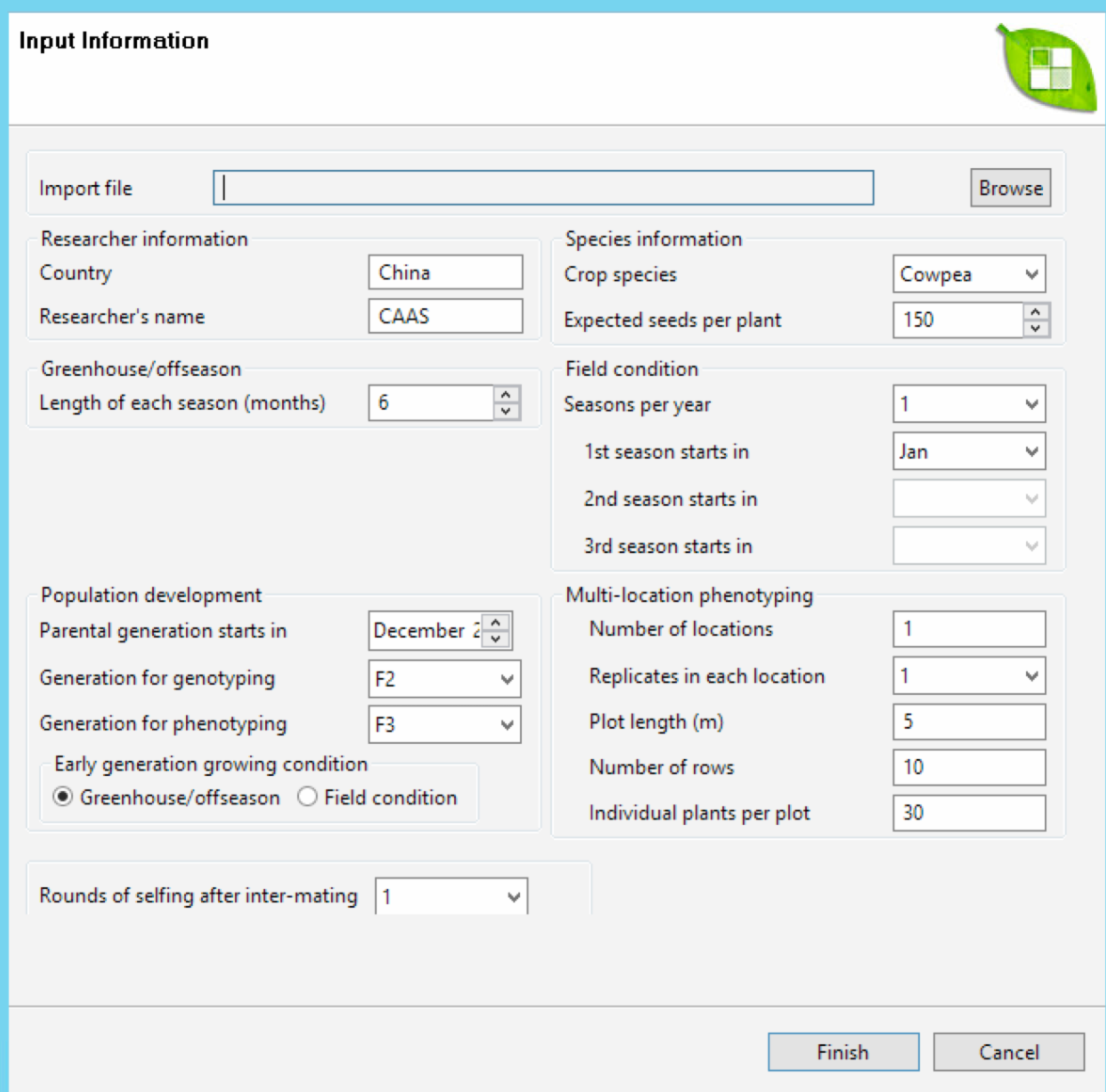
About Expected Seeds Per Plant
- Expected Seeds Per Plant is set to a crop specific default. The user can modify this number, but the input must fall into the min-max range for the selected crop. If input number is smaller than the minimum number, the minimum number will be assumed. If the input number is greater than the maximum number, the maximum number will be assumed.
- Expected seeds per plant is used to determine seed availability for phenotyping. If seed inventory is too low , additional seed increase by selfing will be requested.
- The number of seeds required for phenotyping is calculated from settings for Multi-Locational Phenotyping. For example, multi-locational phenotyping is only possible when there are enough F2 plants to produce the expected amount of seed. Otherwise, phenotyping will be delayed until the required seeds are produced.
Select Finish to populate the parameters and breeding scheme.
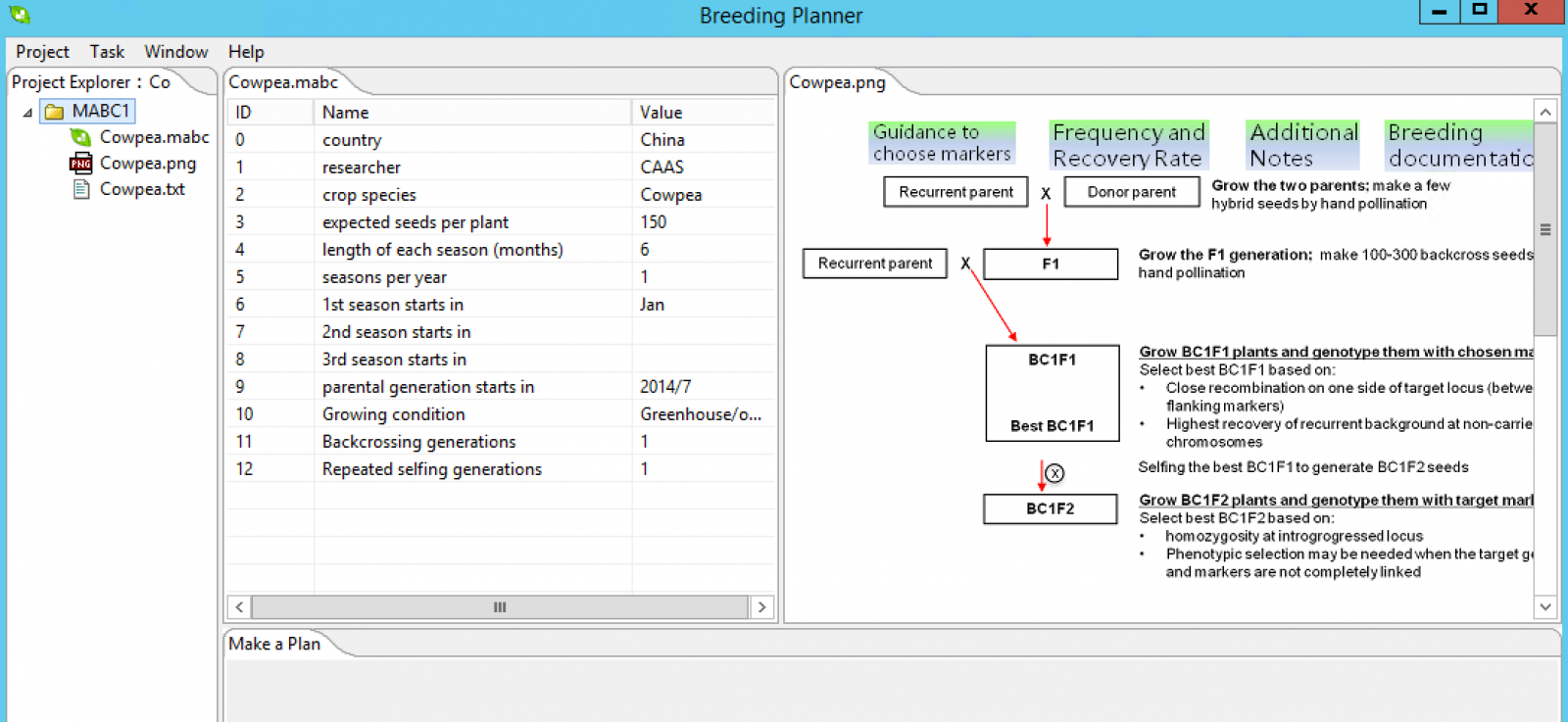
Make a Breeding Plan
Once the parameters are set, right click the breeding method folder, and select Make a Plan from the dropdown menu.
Marker-Assisted Recurrent Selection (MARS) Plan
The marker-assisted recurrent selection (MARS) planning tool schedules events, models QTL to include as selection criteria, proposes selections, plans crosses, and predicts homozygosity in fixed line development.
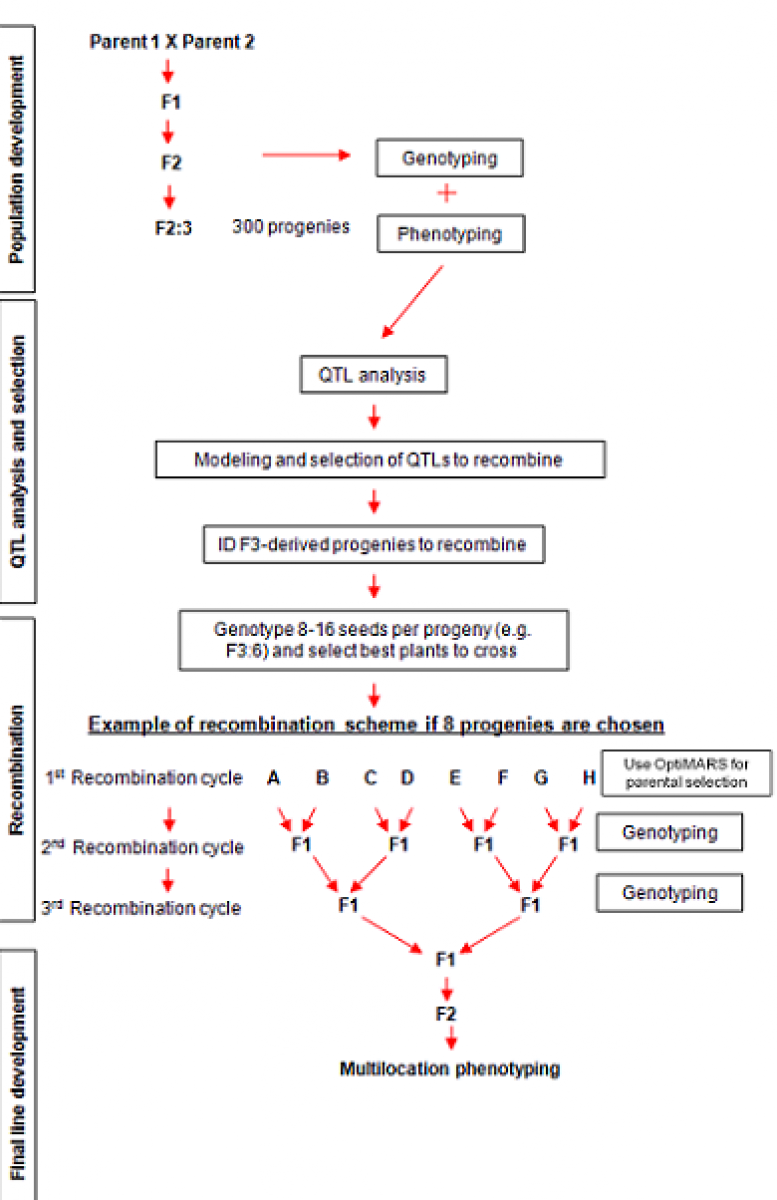
Diagram of a MARS breeding program in cowpea, a self-pollinated species
MARS consists of four main stages:
- Training population development: A training population is evaluated for phenotype in field trials and genotyped using mapped genetic markers. The MARS planning tool will assist in scheduling training population events to meet anticipated deadlines.
- QTL analysis and selection: QTL analysis correlates phenotypic variance to genotypic variance, and identifies putative genetic regions involved in quantitative traits. The MARS planning tool will schedule QTL analysis and model a selection of QTL that optimize the traits of interest for recombination. QTL analyses can be performed with the Breeding Management System’s Breeding View statistical software (see section 5.8) or with other external software applications.
- Recombination: After QTL are identified another Breeding Management Tool, OptiMAS, can be used to identify progeny from the training population for inter-mating. The MARS Planning Tool develops a plan for crossing progeny selected using OptiMAS.
- Fixed line development: The MARS Planning Tool will propose optimum selection scheme for each generation of selfing and predict the homozygosity of the breeding population.
Choose start date and the set the current status of the breeding program. Select the Make a Plan button.
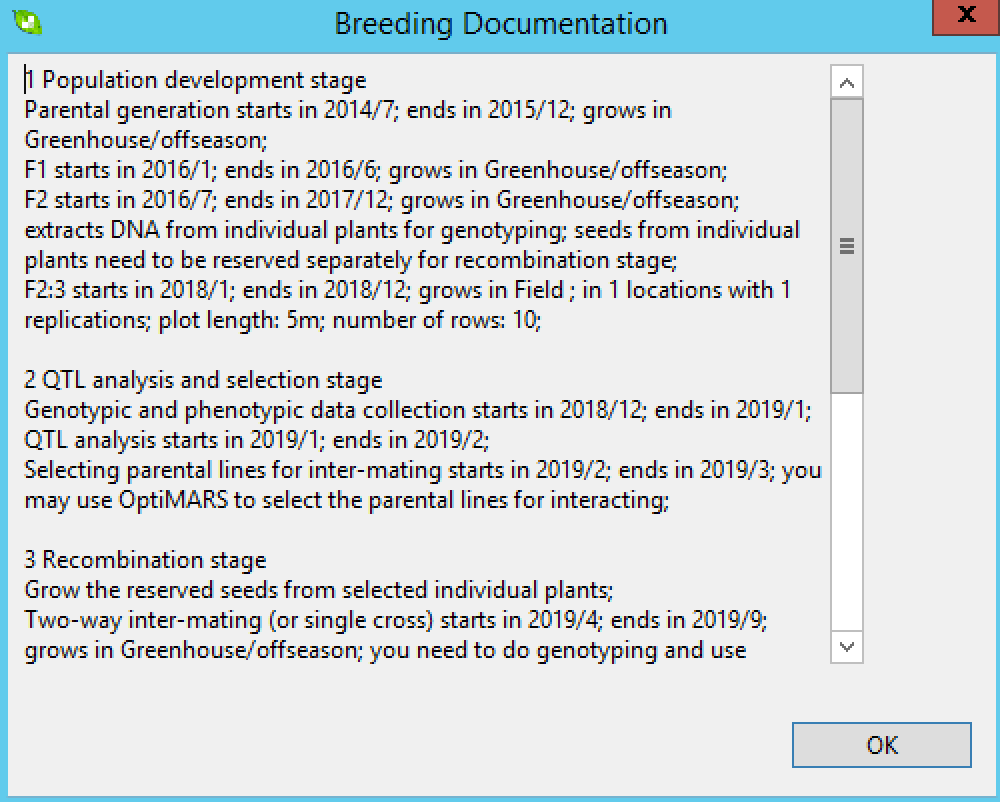
?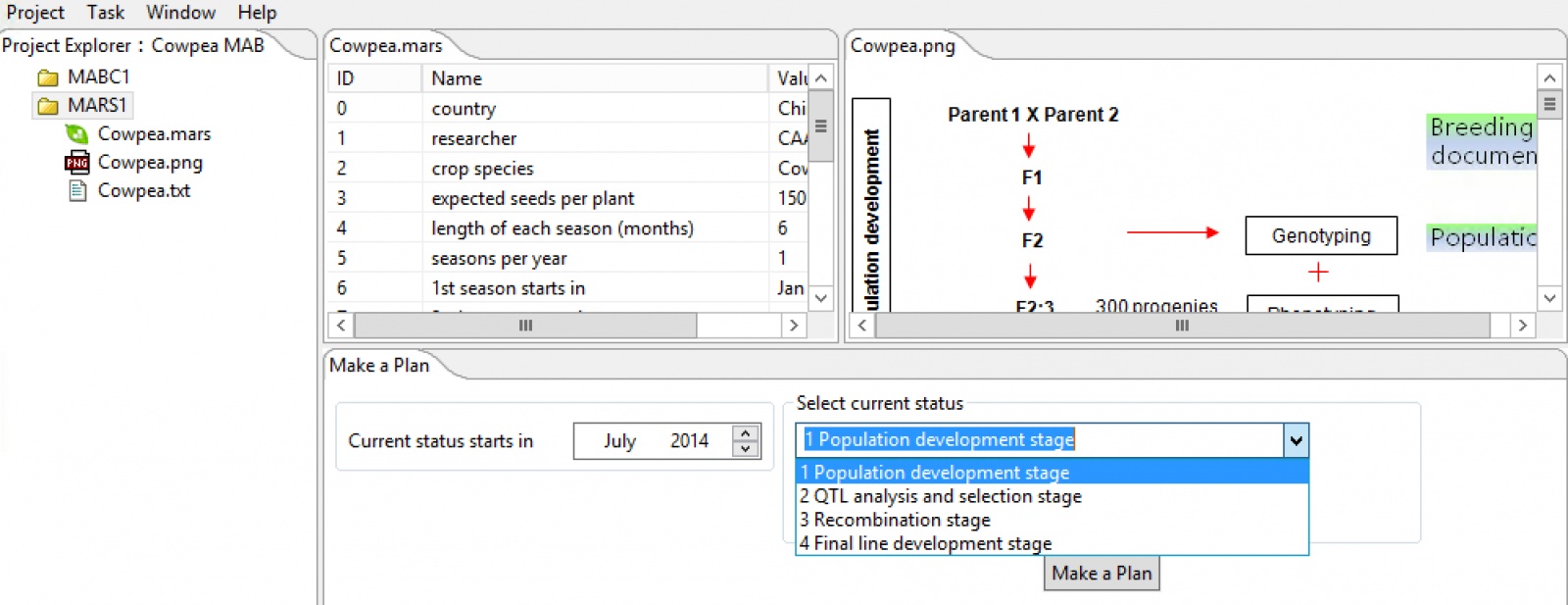
?The breeding documentation window will appear with a timeline for the project accessible as a .txt file in the project folder.
Marker-Assisted Backcross Breeding (MABC) Plan
Choose start date and the current status of the breeding program. Select the Make a Plan button.
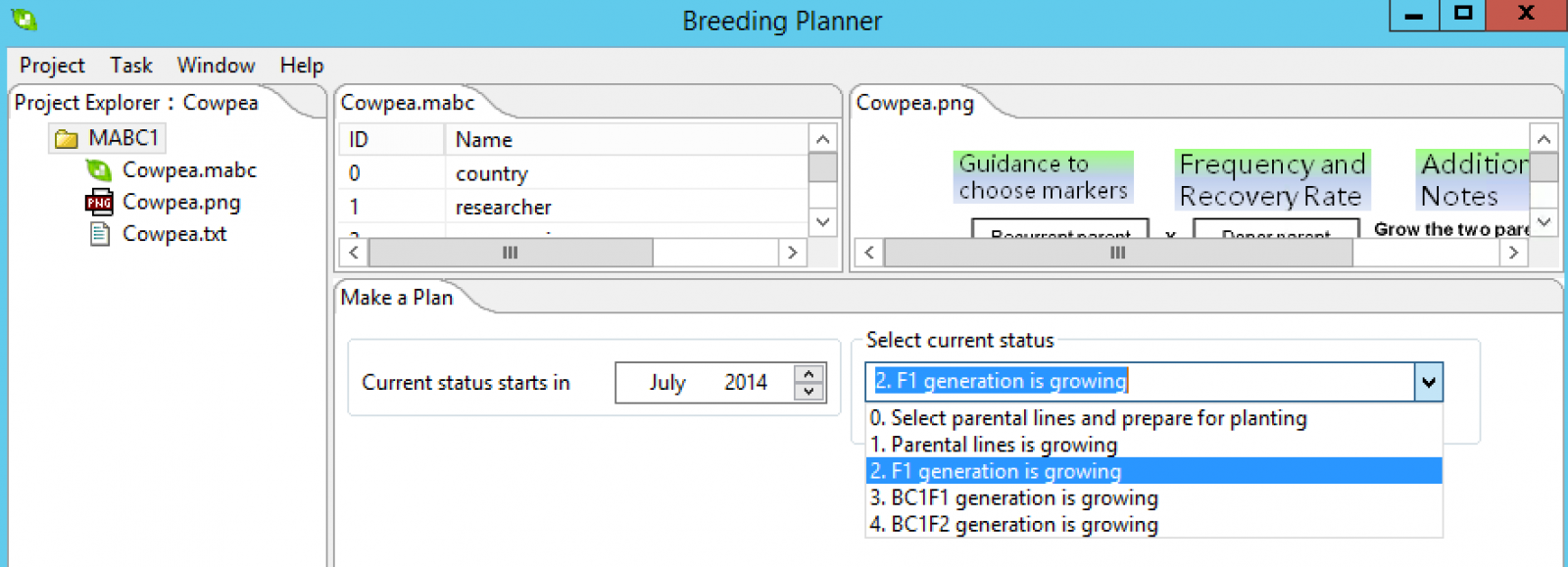
The breeding documentation window will appear with a timeline for the project accessible as a .txt file in the project folder.
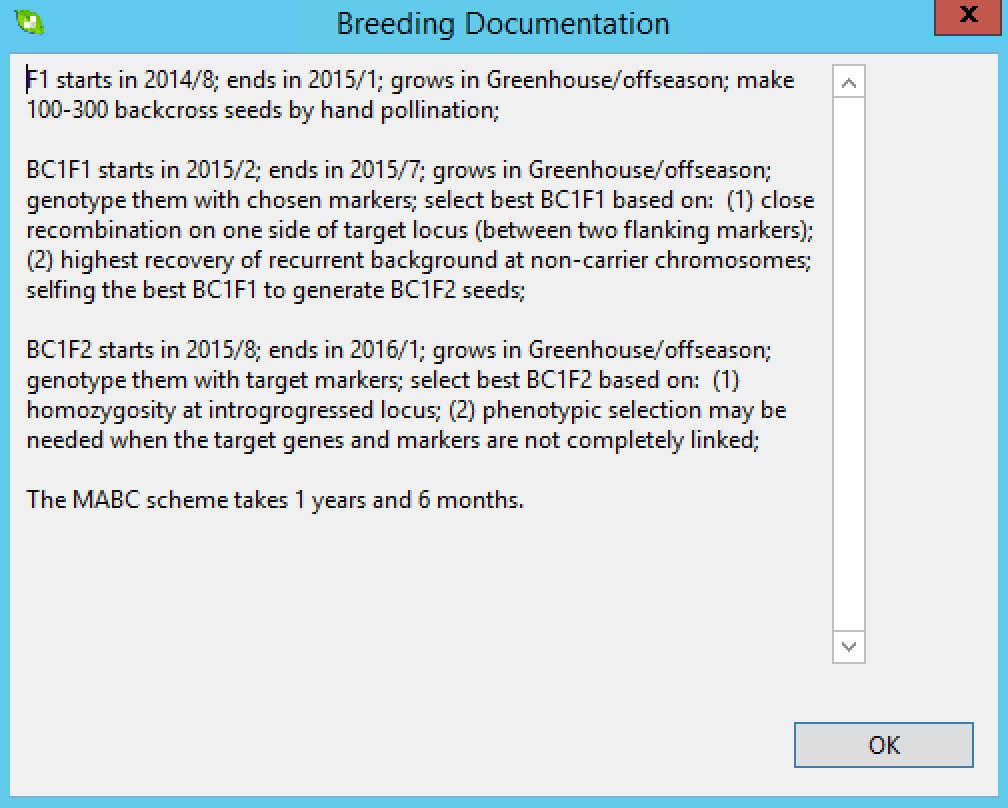
Marker-Assisted Selection (MAS) for Gene Pyramiding Plan
Selection for two or more genes at a time, or gene pyramiding, can be greatly enhanced by marker-assisted section (MAS). Gene pyramiding requires a complex design strategy that outlines the number of markers required, best crossing strategies, and the level of inbreeding to maximize the efficiency of marker implementation. The Molecular Breeding Planner does a simulation analysis to develop breeding plans for gene pyramiding.
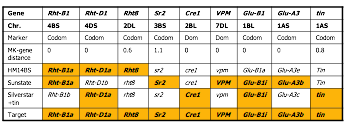
Nine major genes and their distribution in three wheat parental lines
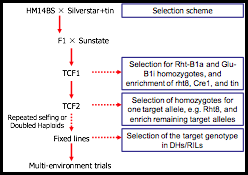
Optimum crossing and selection scheme for gene pyramiding identified by simulation
Save a Project
When the Molecular Breeding Planner is closed, projects are automatically are saved as .ibp files. A folder titled with your project name will appear in the destination folder you established when you created the project. This project folder will automatically be populated with the .ibp project file and subfolders relating to different marker-assisted breeding plans.
Individual breeding plan files that are displayed in the Project window can also be saved independently of the project folder by right clicking and selecting the Save As option.
Open Existing Project
From Open in the Project dropdown menu, you can browse for existing .ibp project files.
Alternatively, from New Project in the Project dropdown menu, you can search for existing projects if you know the name and can specify the root directory.