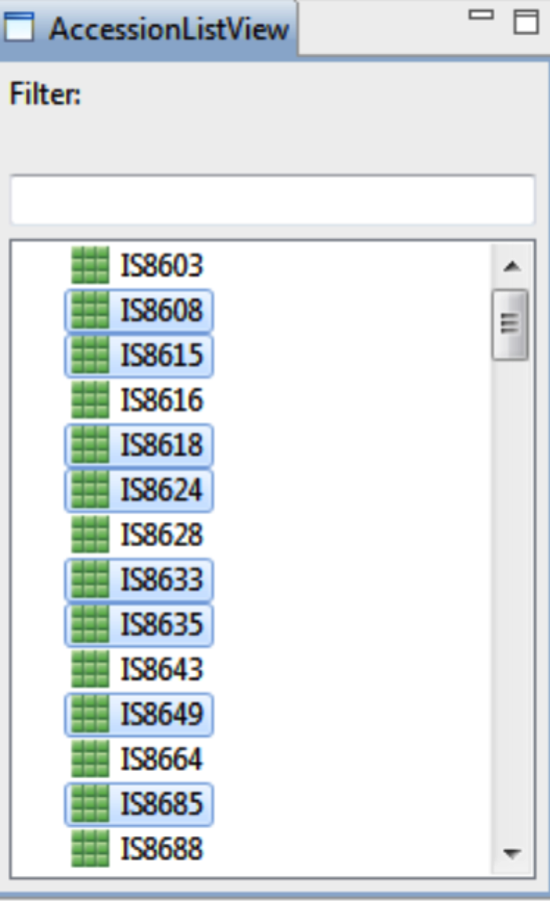
Accession List & Selected Accession View
Select accessions of interest as probable donor or recurrent parents. Use Ctrl/ Shift Key for multiple selections.
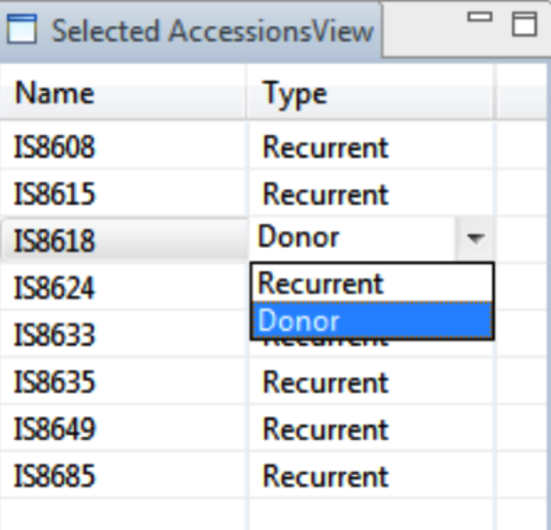
These selections will now appear in the Selected Accessions View. Selections are by default set to Recurrent. Change the default by choosing Donor from the drop down menu.
Sort individuals based on the frequency of alleles for the selected marker. Select marker of interest and click on, or right click on Graphical view. Select Sort.
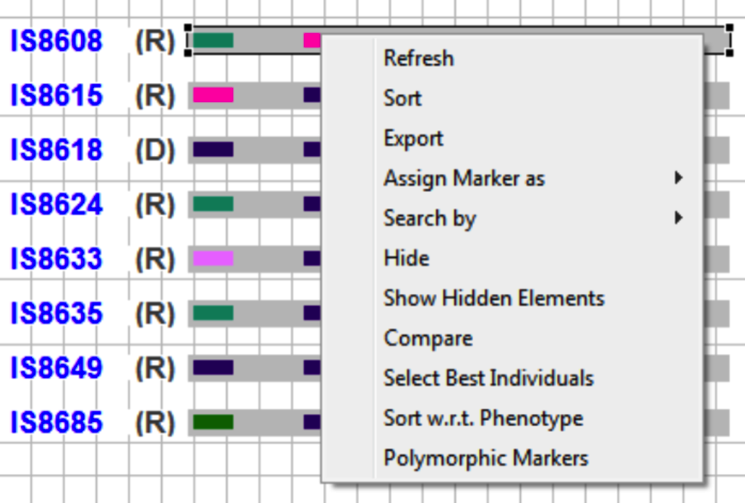

Alternatively, go to the Windows menu option and select Sorting Markers. Choose chromosome and marker, and click Sort.
Sorting displays accession list with rare alleles at top and more common alleles at bottom.

Selected accessions sorted by first marker, umc164c, on chromosome 1
Hide Accessions
Hide the heat maps of selected accessions in Graphical View. Select the appropriate heat maps. Right click on selections and click Hide.
Reveal Hidden Accessions
Right click on selection and click Show Hidden Elements. Choose the hidden accessions that you would like to reveal and click Finish.
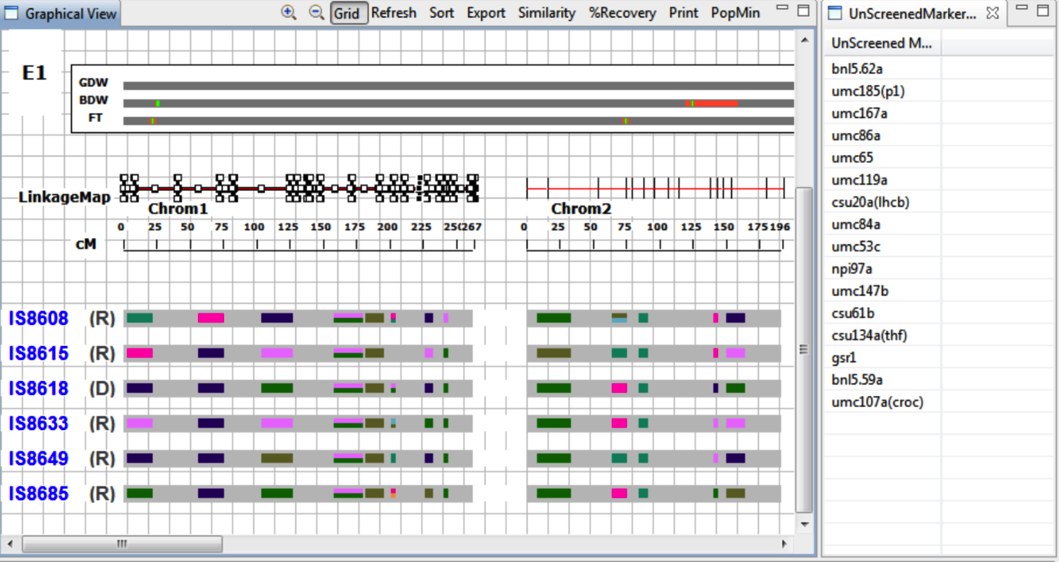
Graphical View
Select Refresh to update changes made in the Selected Accession View. Heat maps of selected accessions display genotype information as coloured boxes allowing comparison of genotypes with linkage and QTL map.
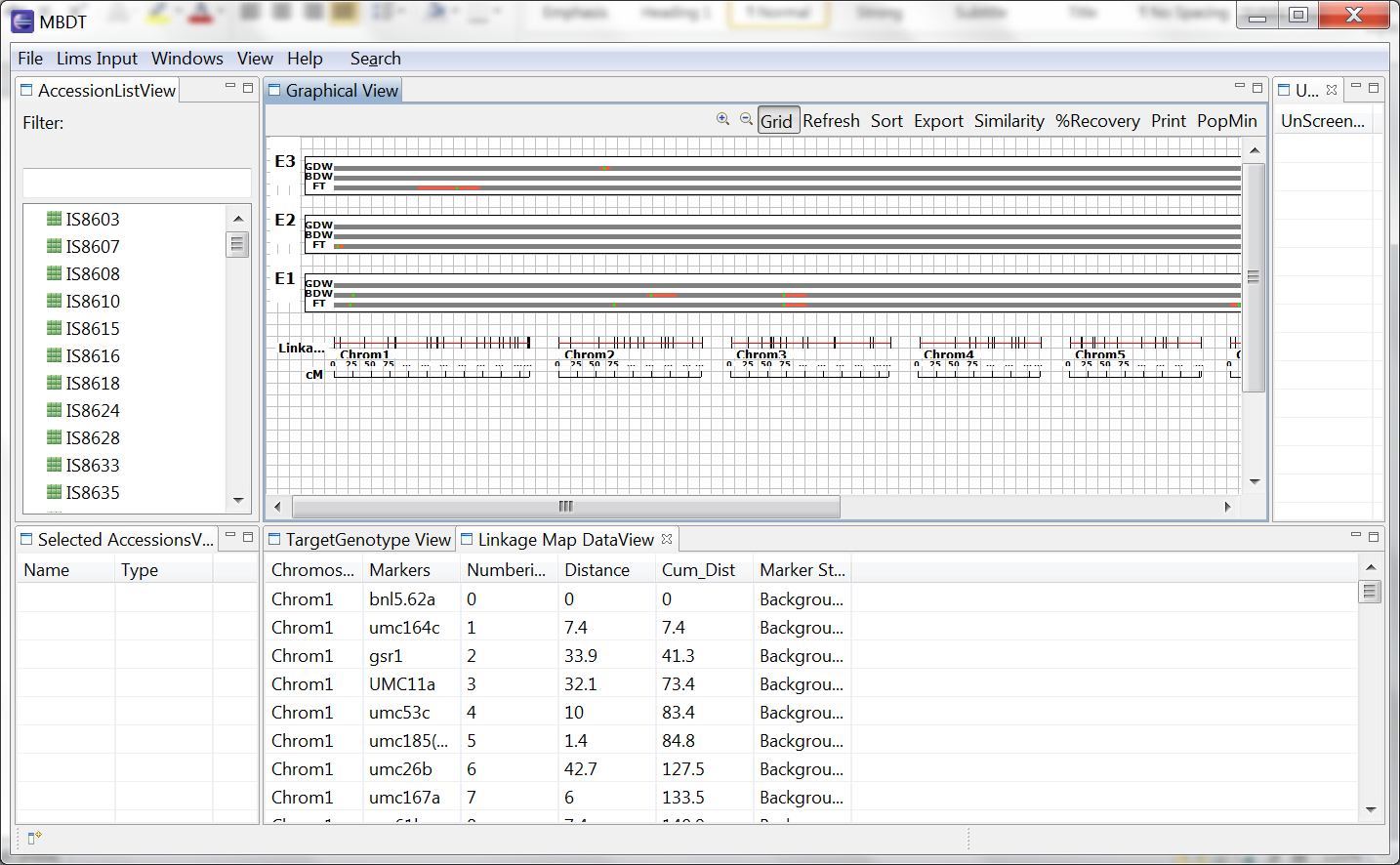
The accessions are now visible from the graphical view.
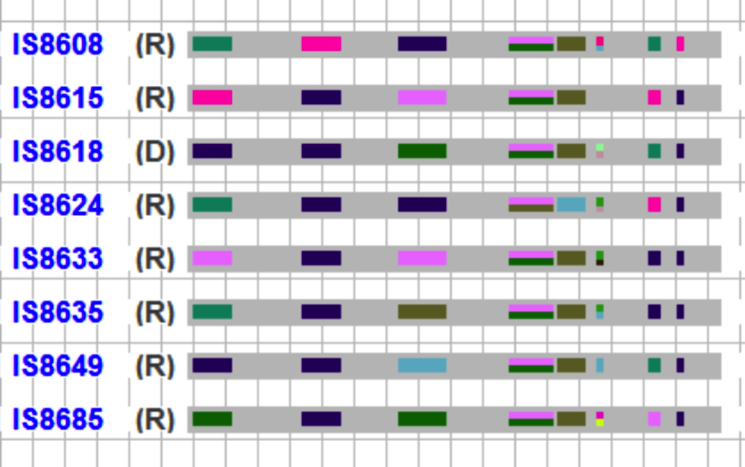

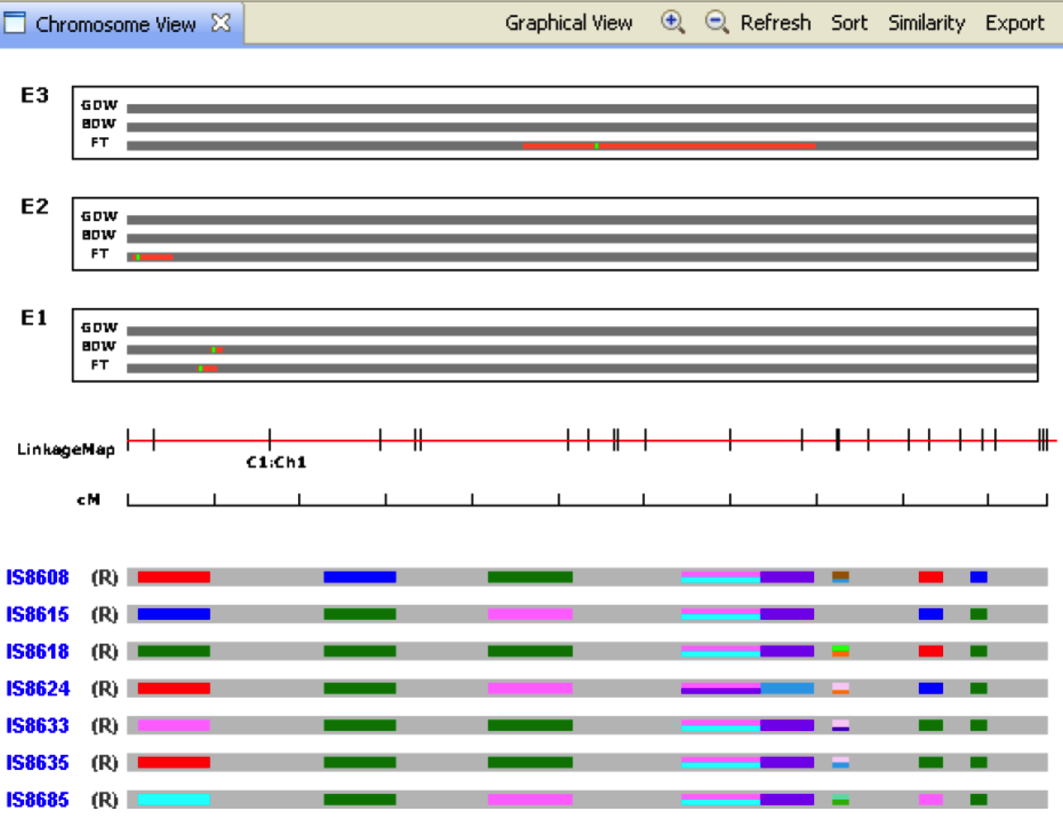
R and D represent recurrent and donor parent assignment. Colored boxed on the grey bar represent the screened marker for which genotype data is available. The markers are coloured based on the allele values. The missing values are coloured in background colour, light grey to represent un-genotyped sections of the chromosome.
Click on QTL region to view QTL information in pop-up window.
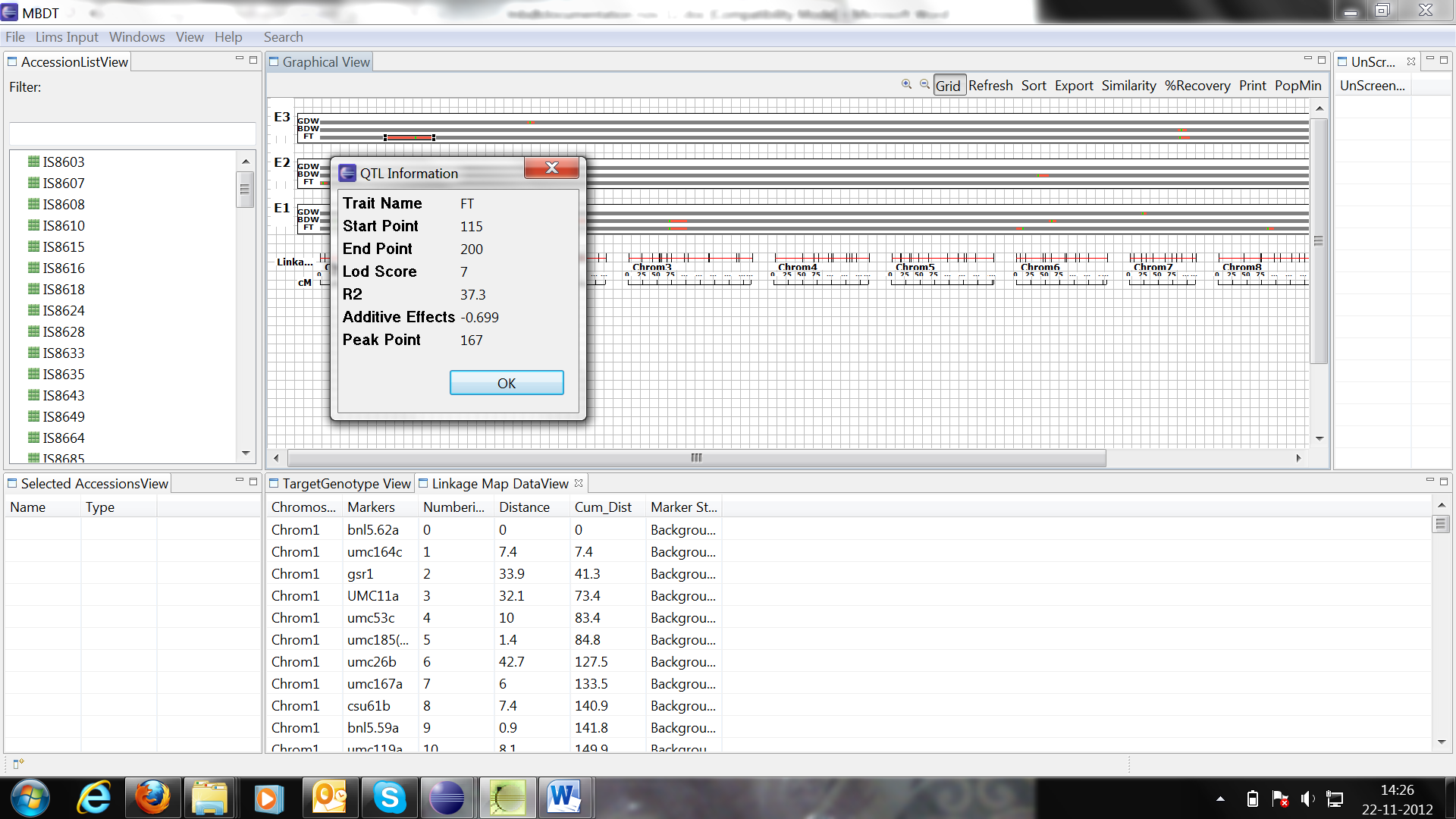
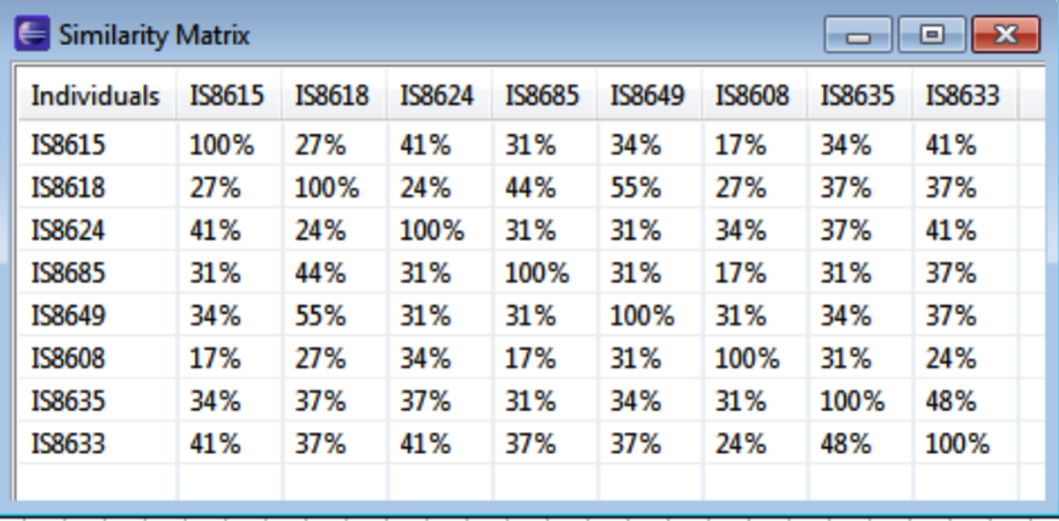
Similarity Matrix
Select Similarity to generate a similarity matrix to visualize the percentage of similarity between selected parents based on the number of shared alleles.
Unscreened Markers
Select an area of linkage map to populate the Unscreened Marker view.

The unscreened markers display as a list.
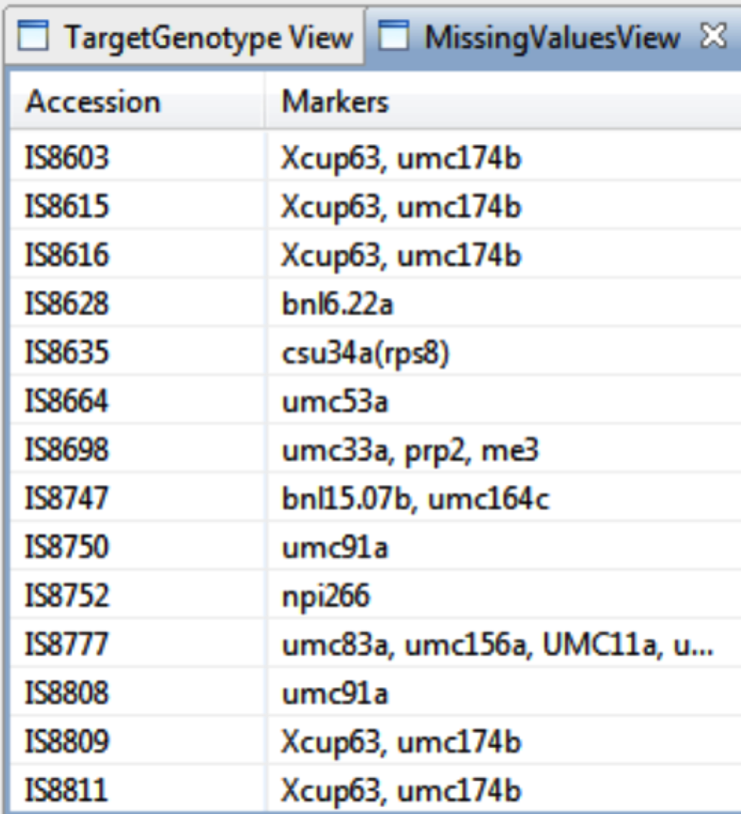
Missing Markers
View the missing markers for each accession.
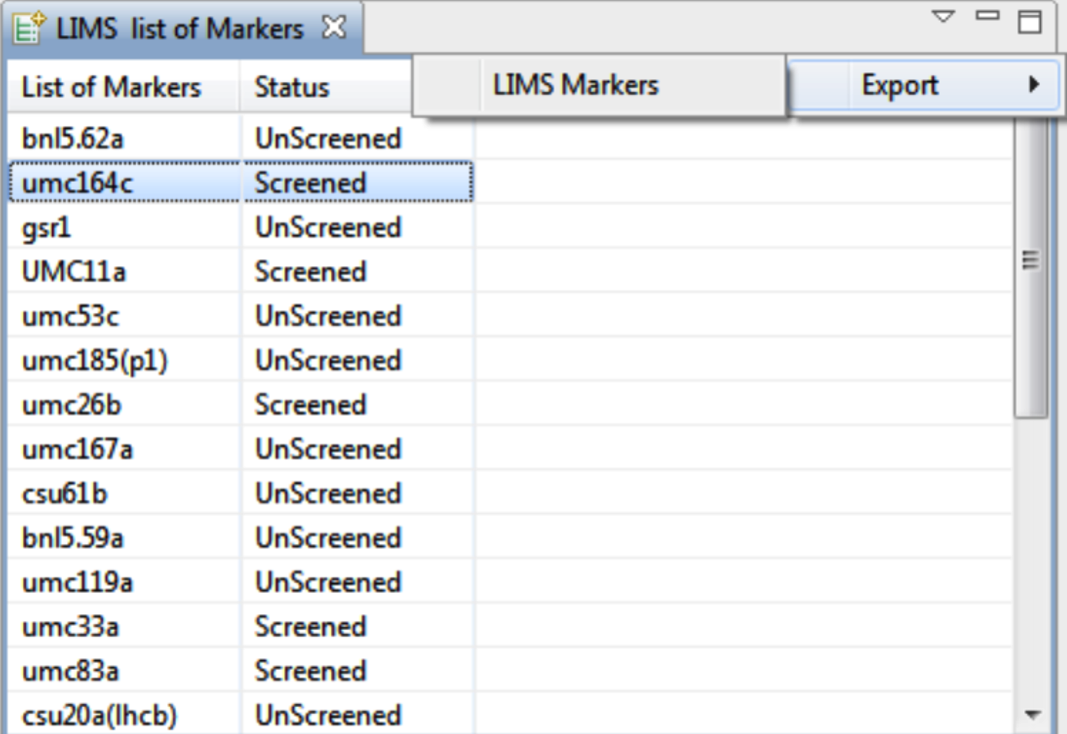
Lims Markers
Identify genotyped markers that match markers within the linkage map. Select Lims Input menu > Lims Markers. Select OK.
Delete markers from the list by right clicking selections and choosing Delete Selected Marker. Select Export > Lims Markers to save this list as a .txt file.
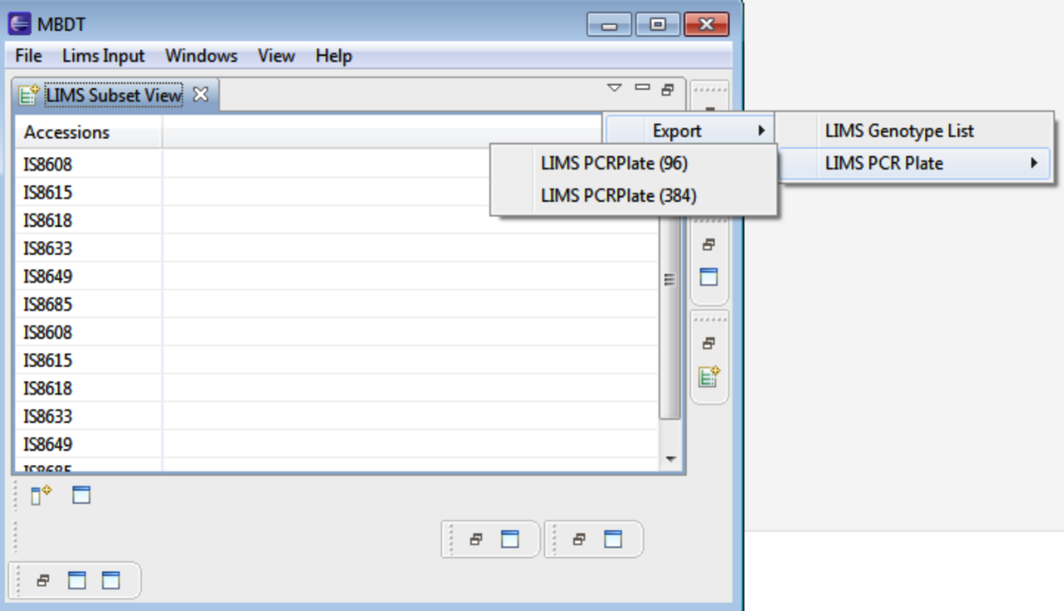
Lims Subset
Select Lims Input > Lims Accessions. The Lims Subset View allows export of the genotype list as well as a file outlining 96 or 384 well PCR plates.
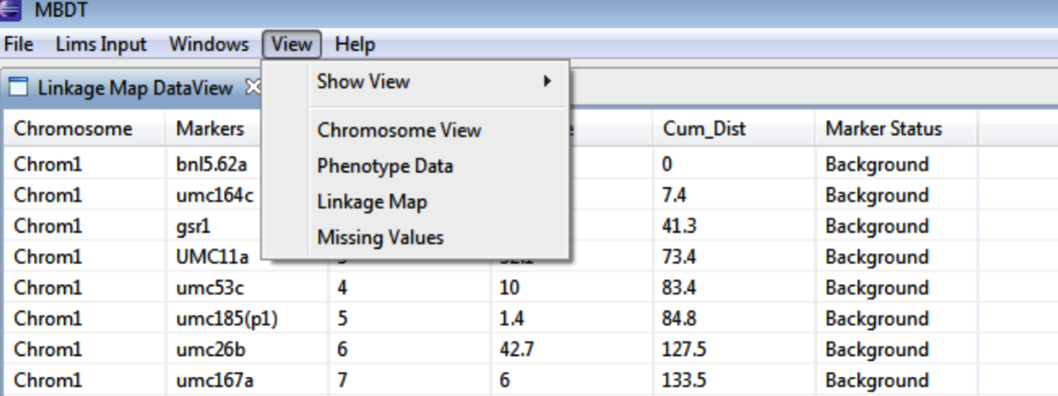
Linkage Map Information
View information about the linkage map by selecting this view.
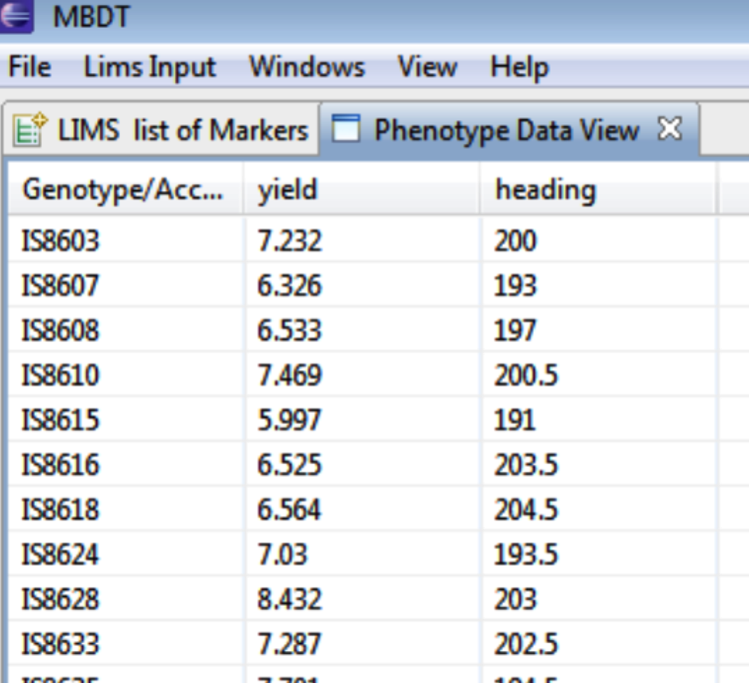
View Phenotypic Data
Sort phenotypic data by double clicking the trait name.
Chromosome View
Limit graphical view to a single chromosome by selecting Chromosome View and choosing a chromosome of interest.
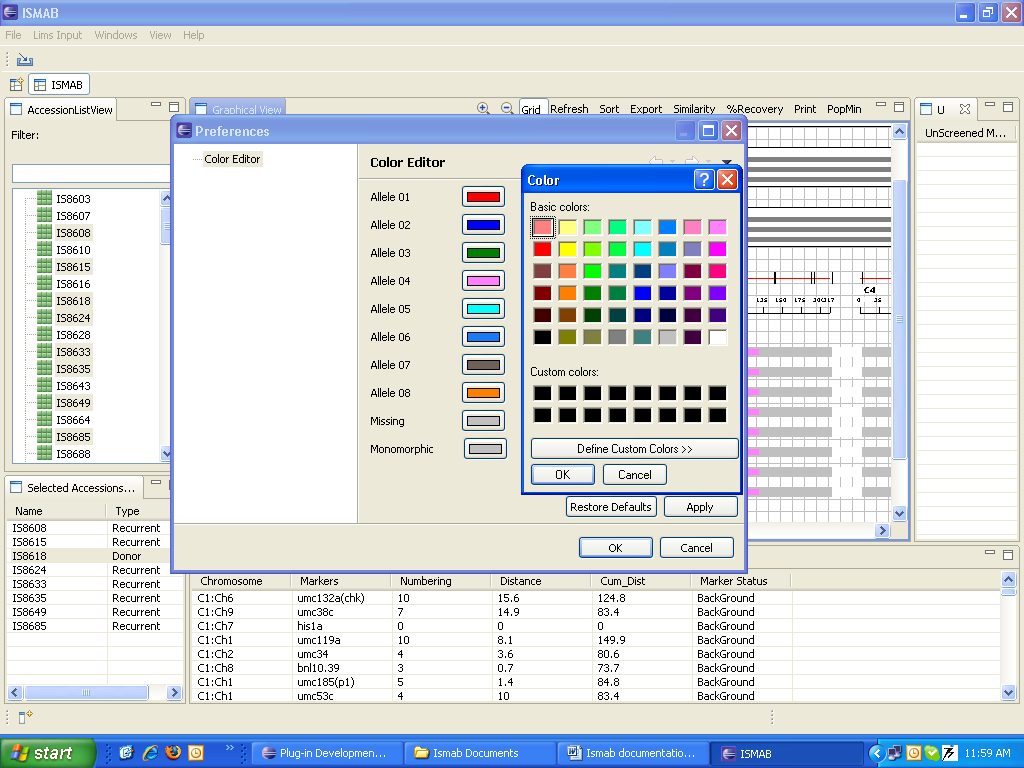
Customize Marker Colors
Change the default colors by select Preferences under the Windows menu. Change the color of any allele using the color editor.