Assign Foreground & Background Markers
Foreground Markers are the markers linked to QTL from the donor parent. Background markers are unique to the recurrent parent and are used to track the recovery of the recurrent parent genome. By default all markers are defined as background markers.
Assign markers as Foreground markers by selecting a QTL on the QTL map or a portion of the linkage map. Right click and choose Assign Marker as > Foreground.
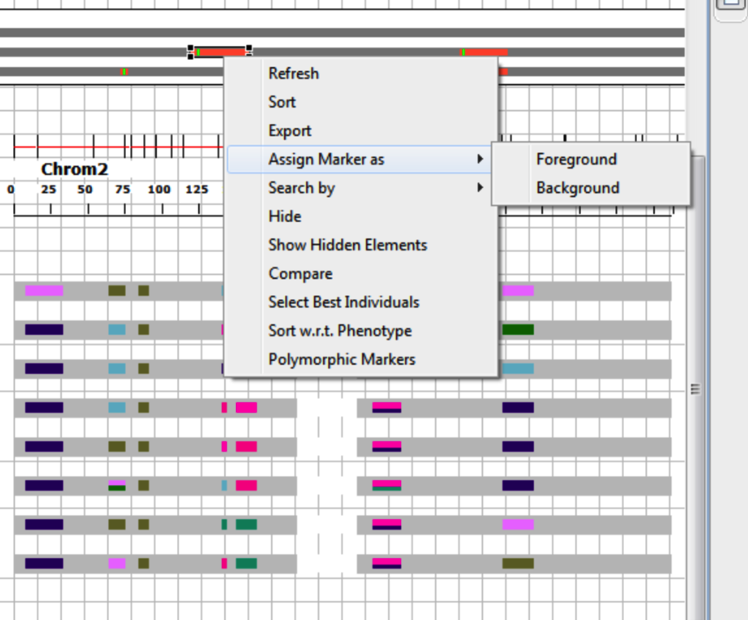
Assigning a foreground marker will result in a broadening the representative rectangular boxes. The status of marker in the LinkageMap data view will be changed accordingly.
Revert foreground markers to background markers by selecting the QTL or a portion of the linkage map. Right click and choose Assign Marker as > Background. The status of marker in the LinkageMap data view will change accordingly.
Find Polymorphic Markers
Compare potential parents based on their polymorphic markers by selecting two accessions and right clicking to selecting Compare from the drop down menu.
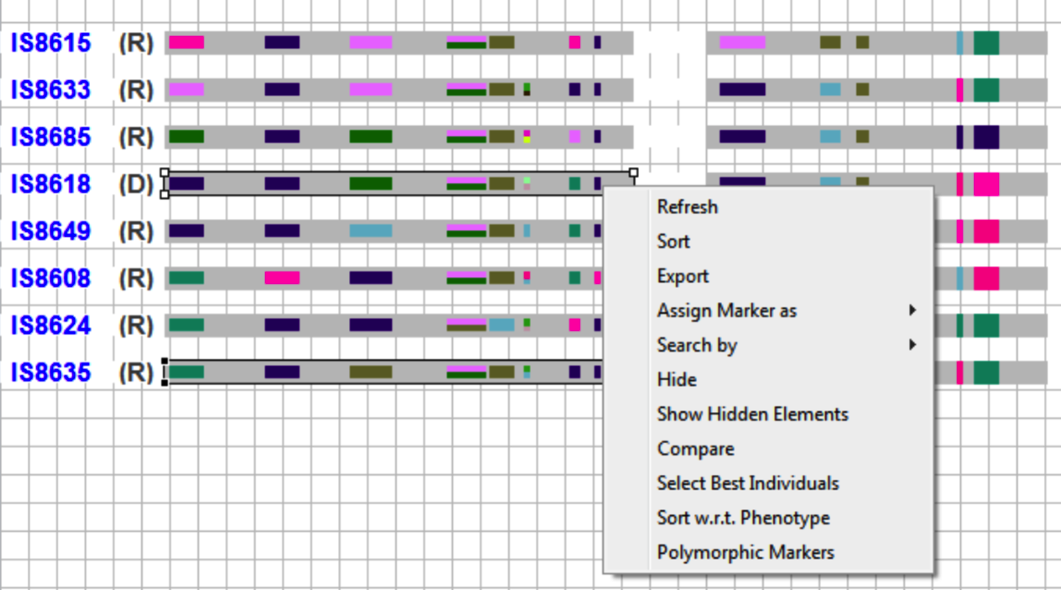
The graphical field will narrow to the two selected accessions.
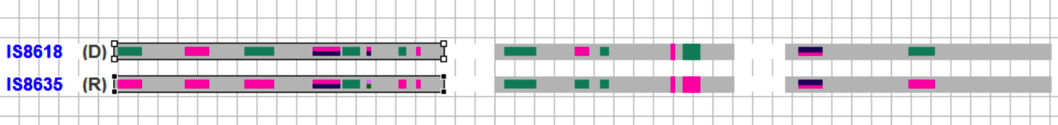
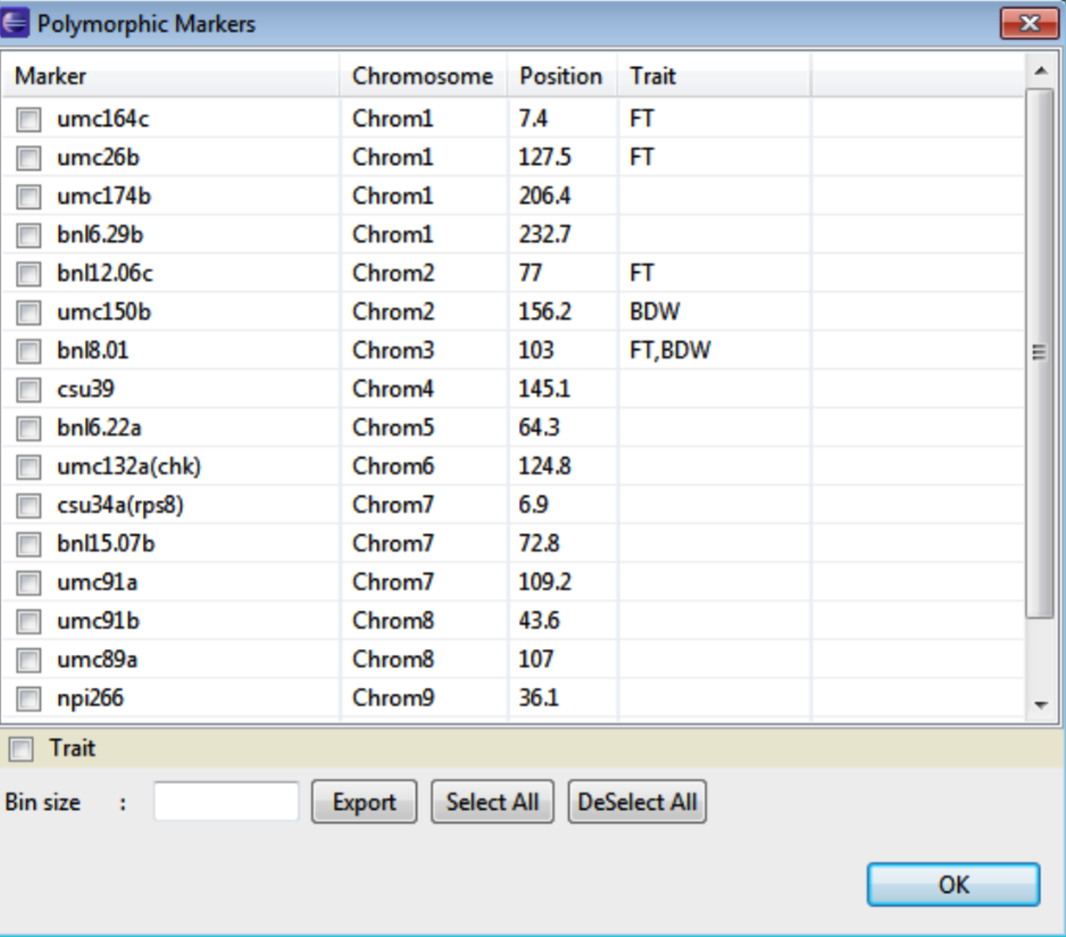
Select both accessions. Right click to choose polymorphic markers from a list.
Create Target Genotype
Select the donor and recurrent parents in graphical view. Select the Target Genotype button from Target Genotype View. The target genotype illustrates polymorphic markers between donor and recurrent parent. Markers in the target genotype represent the recurrent parent.
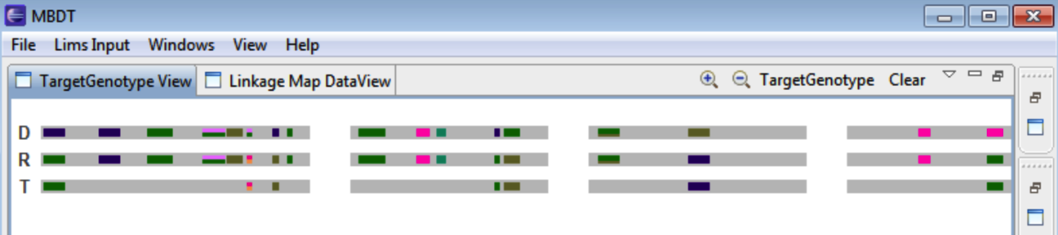 D donor parent (IS86818) and R recurrent parent (IS8685). The T target represents the markers that differ between the parents
D donor parent (IS86818) and R recurrent parent (IS8685). The T target represents the markers that differ between the parents
Replace recurrent parent marker alleles with the desired forward markers from the donor parent. Select the donor parent marker and drag and drop to the target marker location.
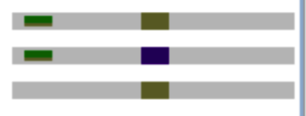
Chromosome 3: Donor parent marker (top) dropped on the target genotype (bottom)
Save the movements of markers while preparing Target Genotype by clicking on drop down arrow and choosing Action and then History of movements. A click on History of movements will open a Save As File Dialog to save the list of moved markers into text file with *.txt file.
Determine Population Size for Next Generation
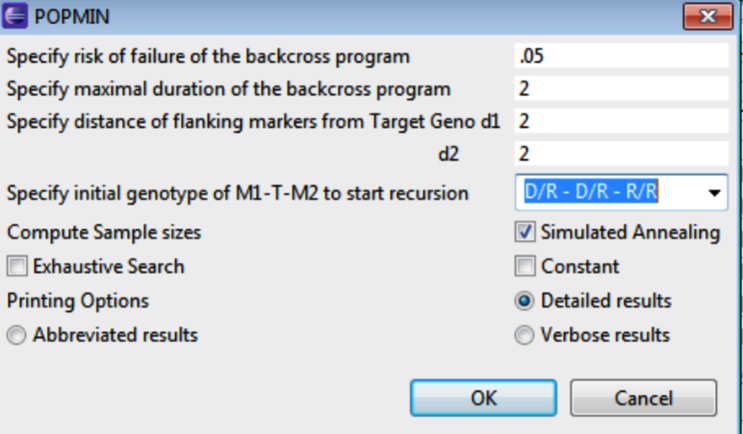
Determine the minimum number of individuals that must be genotyped in successive backcross generations. The PopMIN tool estimates the population size required to obtain at least one double homozygote for the two markers flanking target QTL. Select PopMIN, enter the appropriate values and click OK.
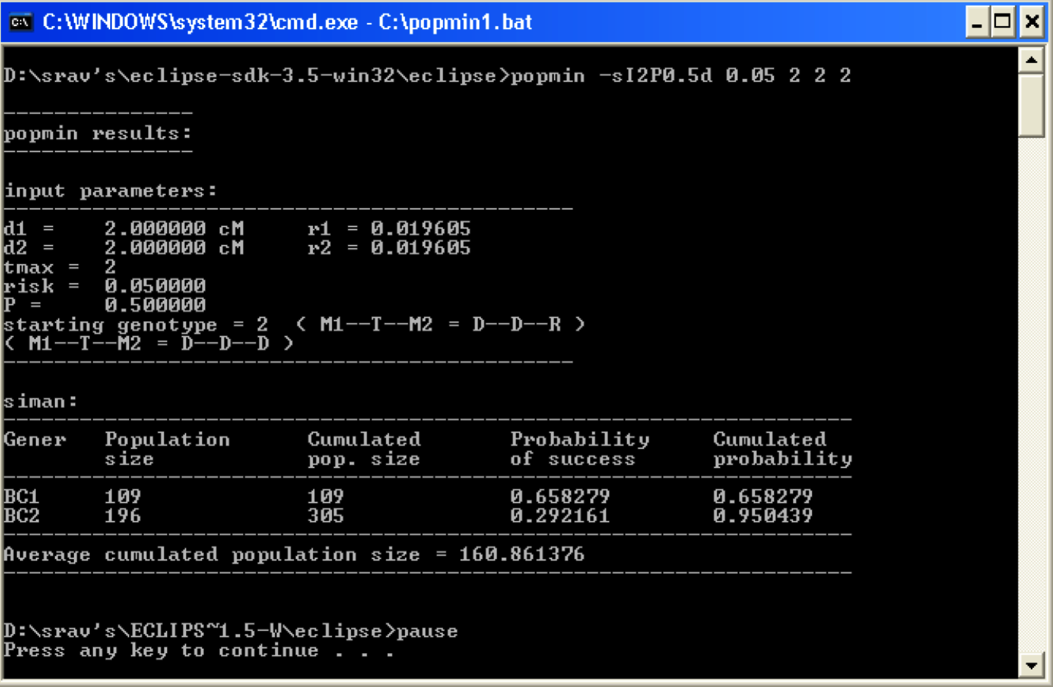
PopMIN results are displayed in the terminal.
Advanced Generations
As subsequent generations are genotyped they can also be viewed with Molecular Breeding Design Tool. Select File > Load Target Data. Select the project and the previous generation from the drop down menu. Load the current generations genotype data, and click Finish.
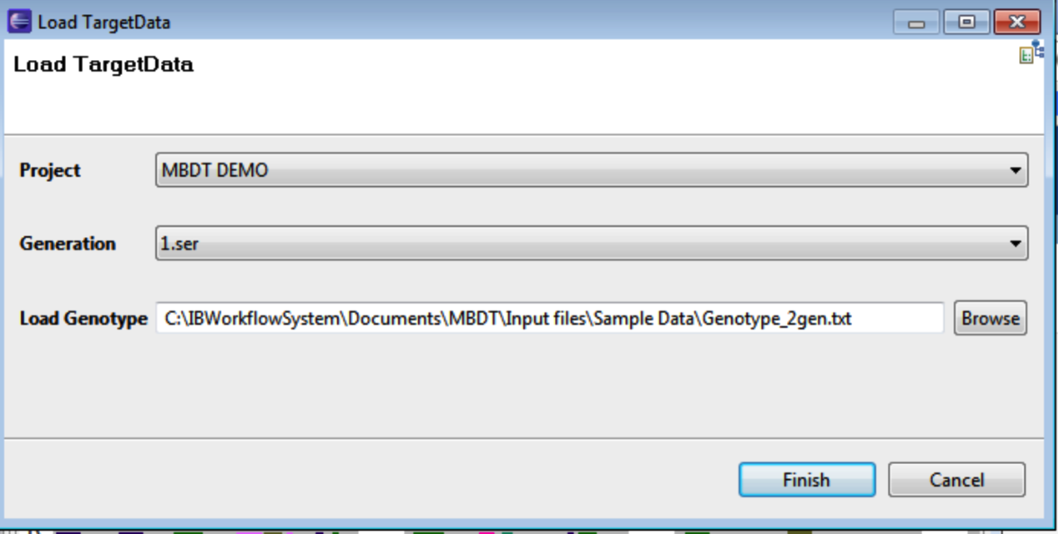
The second generations genotype is graphically displayed beneath the previously established target genotype.
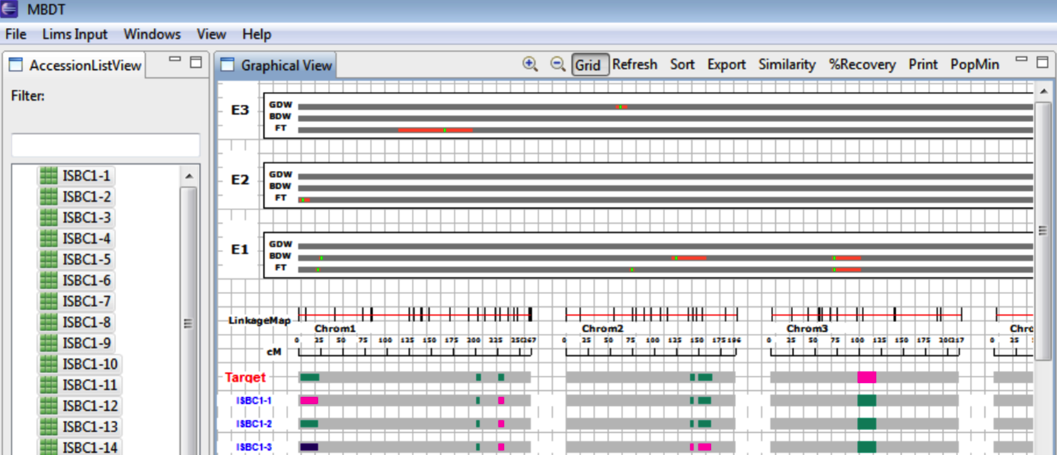
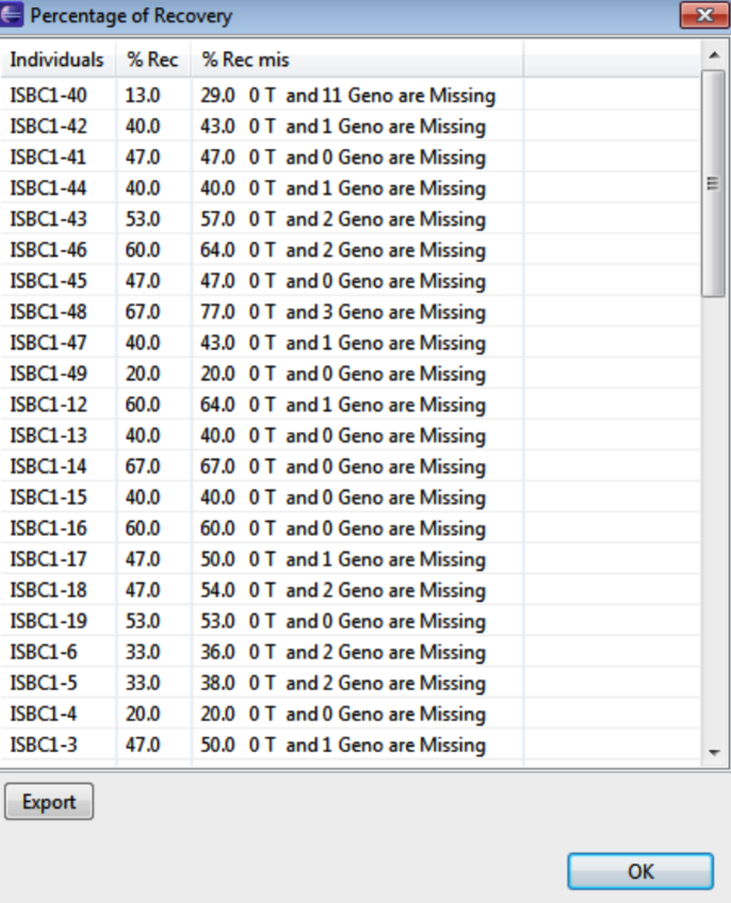
Percentage Recovery
Select Percentage of Recovery to view the proportion of recurrent parent markers in the second generation accessions.
Select Best Individuals
Select Accessions of interest and right click to select the Best Individuals. Choose the foreground marker(s) to base the selection. Include missing information by selecting Missing. Click Finish.
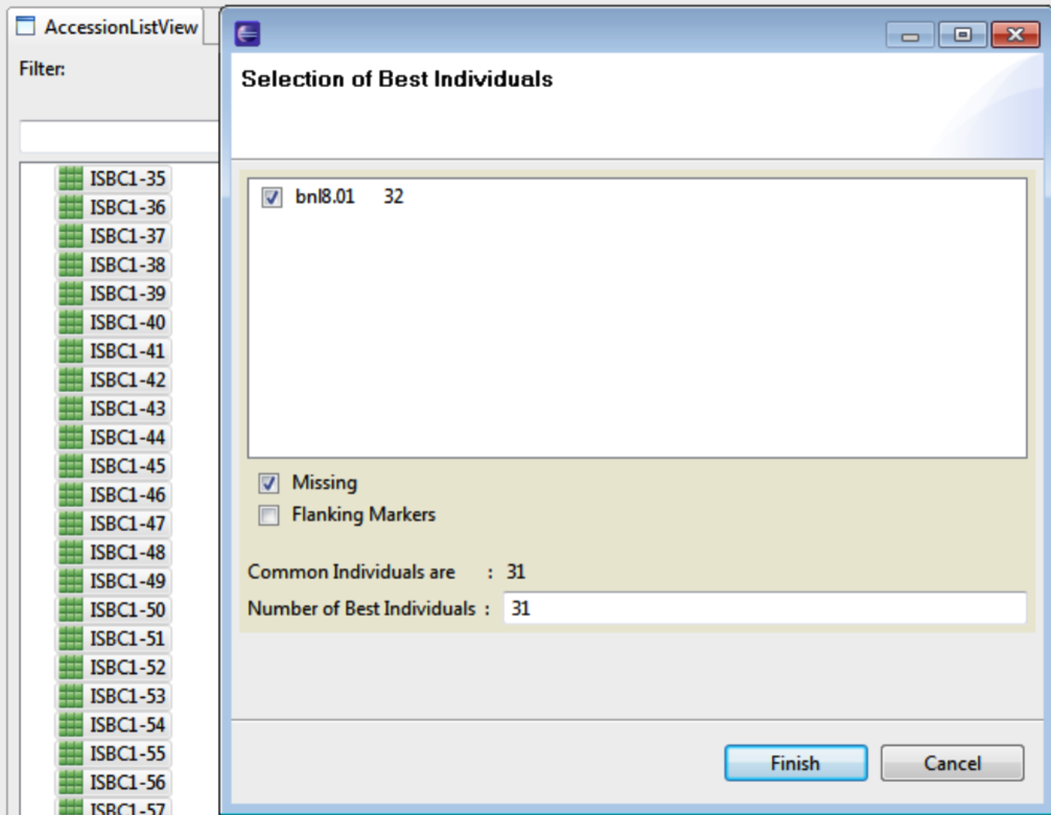
The best individuals are highlighted in the Access List View.