Summary
This tutorial describes how to design a field trial to evaluate a set of germplasm with 32 entries for 6 phenotypic traits in a replicated field trial with three replications in 4 different locations. The experimental design is a resolvable incomplete block design, specifically an alpha lattice design.
- Design Trial
- Create Planting Labels (Optional)
- Reserve & Withdraw Trial Germplasm: Optional
- Related Materials
Design Trial
Trial Details & Settings
- From Manage Studies tool, select Start a New Study.

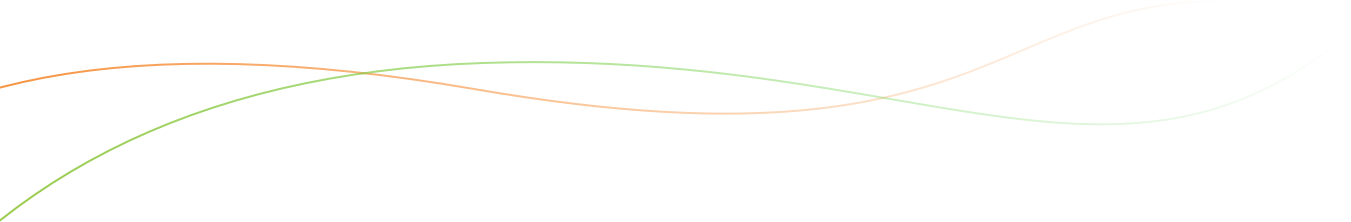
- Select Trial as the Study type and establish basic trial details.
- Name: Performance Trial
- Description: 2018
- Objective: Preliminary yield trial
- Select Add in Study Settings to add management details. Select STUDY_INSTITUTE and PI_NAME_TEXT from dropdown menu. Fill with your details. Select Save.
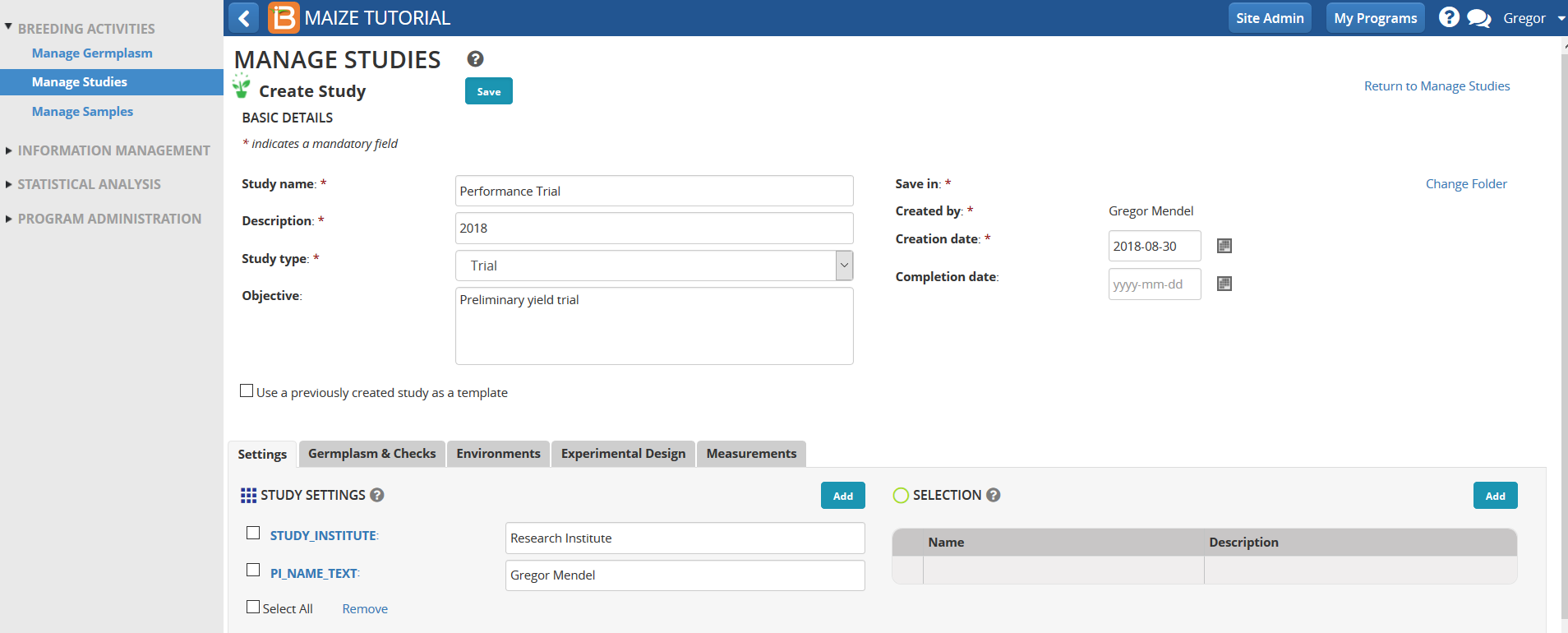
- Select the "+ icon". Enter 2018 as folder name. Select the green check icon after highlighting the Studies folder. The folder named 2018 is created within Studies.
- Select to complete saving the trial.
Germplasm
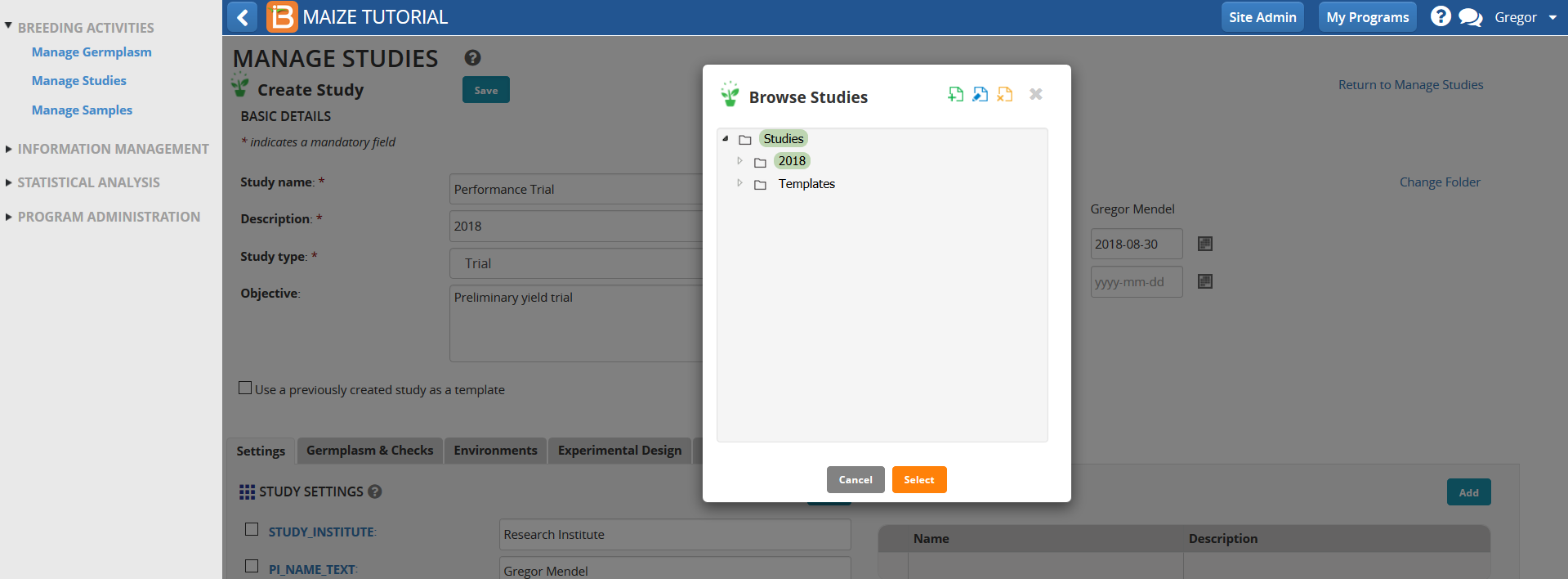
- Under Germplasm Tab, Browse for a germplasm list.
- Select the Trial Germplasm 2018 list.
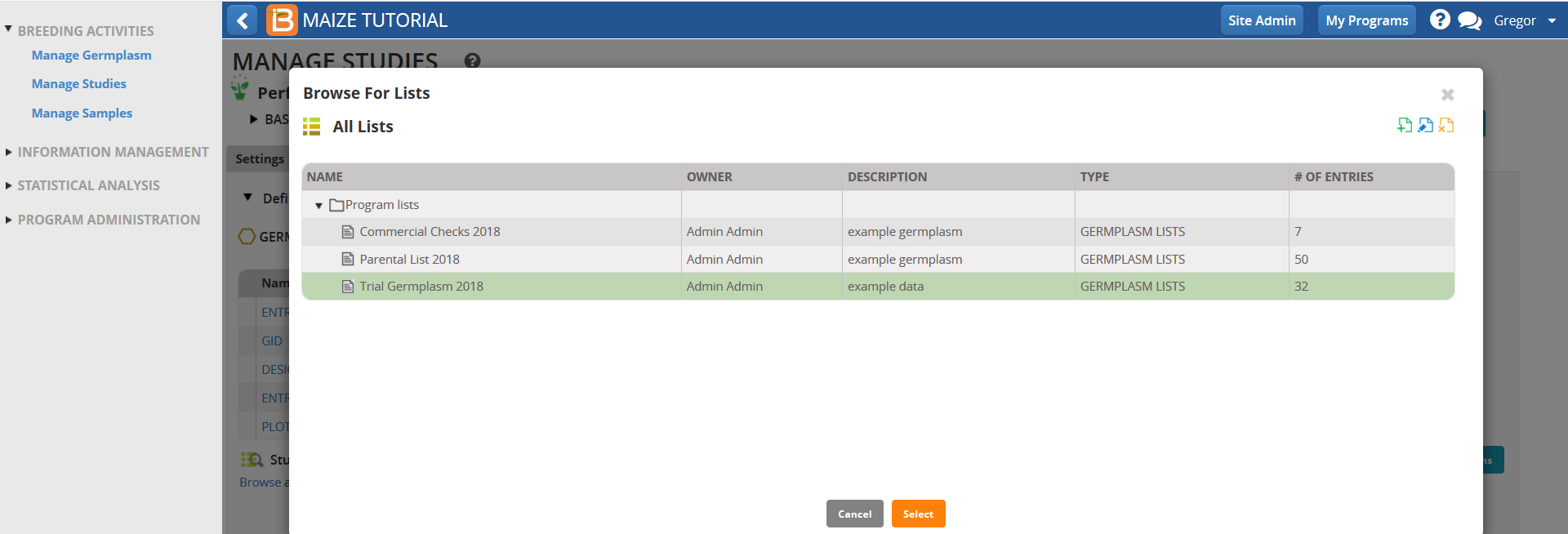
The 32 entry trial germplasm list is now loaded into the performance trial.
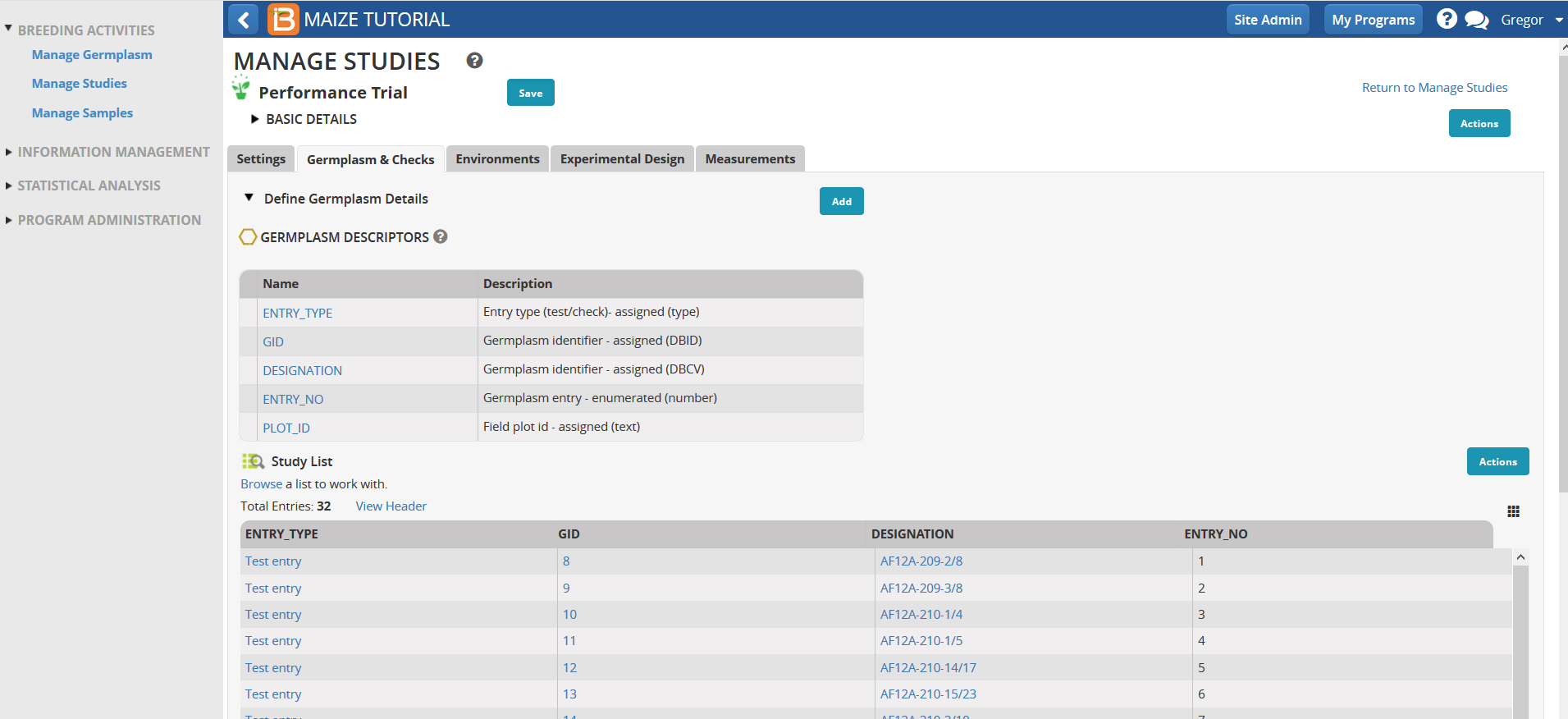
Environments
- Under Environments Tab, specify four environments for the study and select OK. Four environments will now populate the screen below.
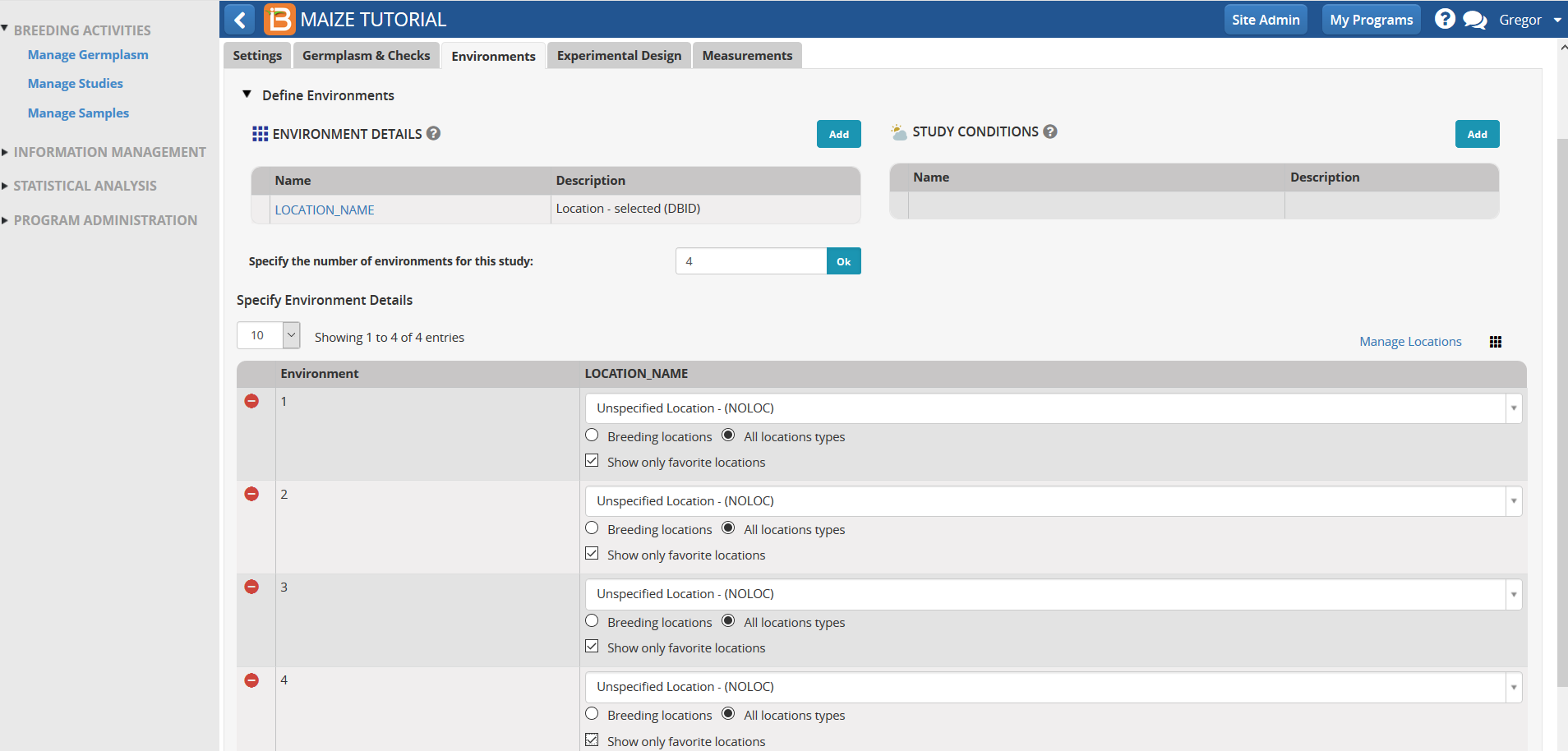
The four environments associated with the tutorial can be found by highlighting the box next to 'Show only favorite locations'.
The 4 environments correspond to the following location names:
- Environment 1: Agua Fria - (AF)
- Environment 2: Sabana Del Medio - (SDM)
- Environment 3: Jutiapa - (JUT)
- Environment 4: Tlaltizapan - (TLA)
- Select the correct location name from the favorite location drop down menu. The correct name should match the environment number of the example data.
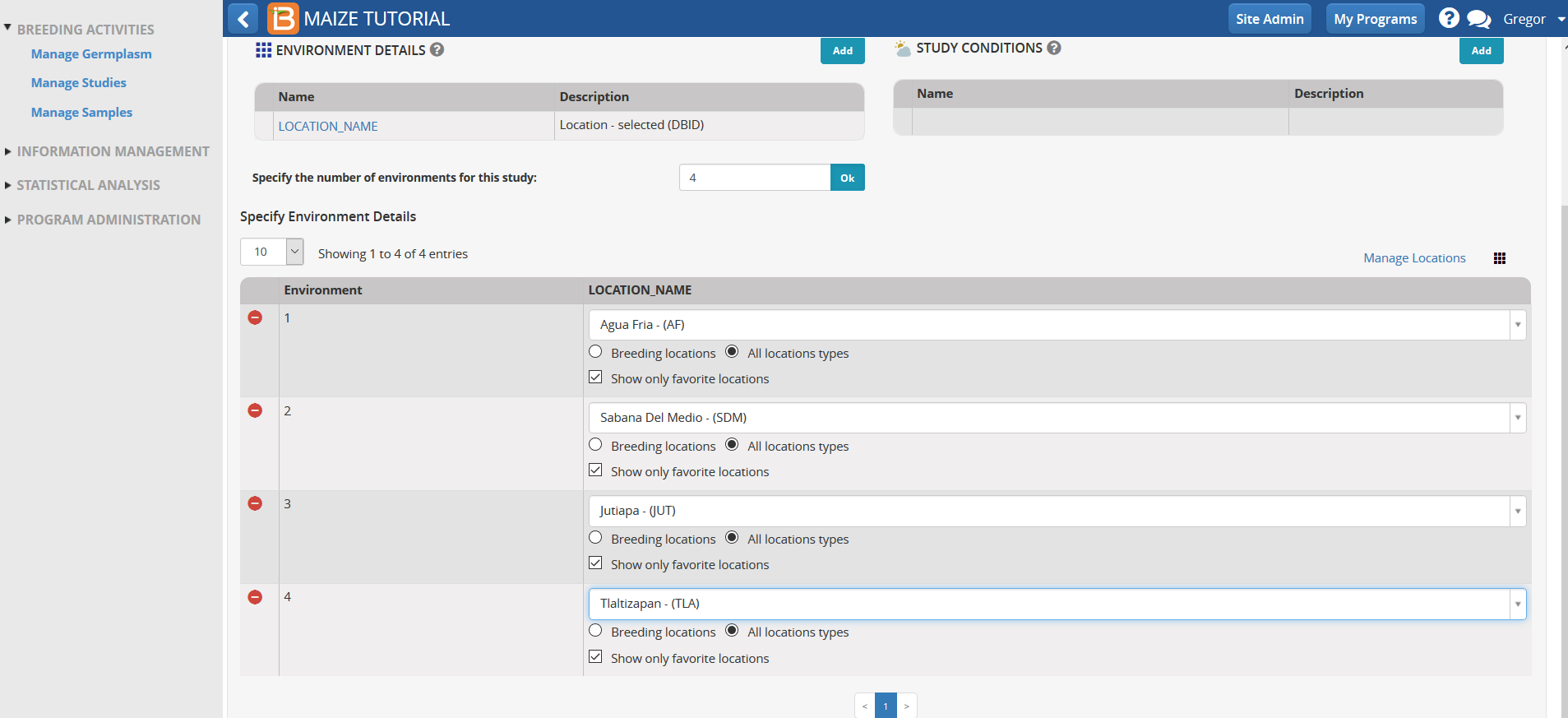
Experimental Design
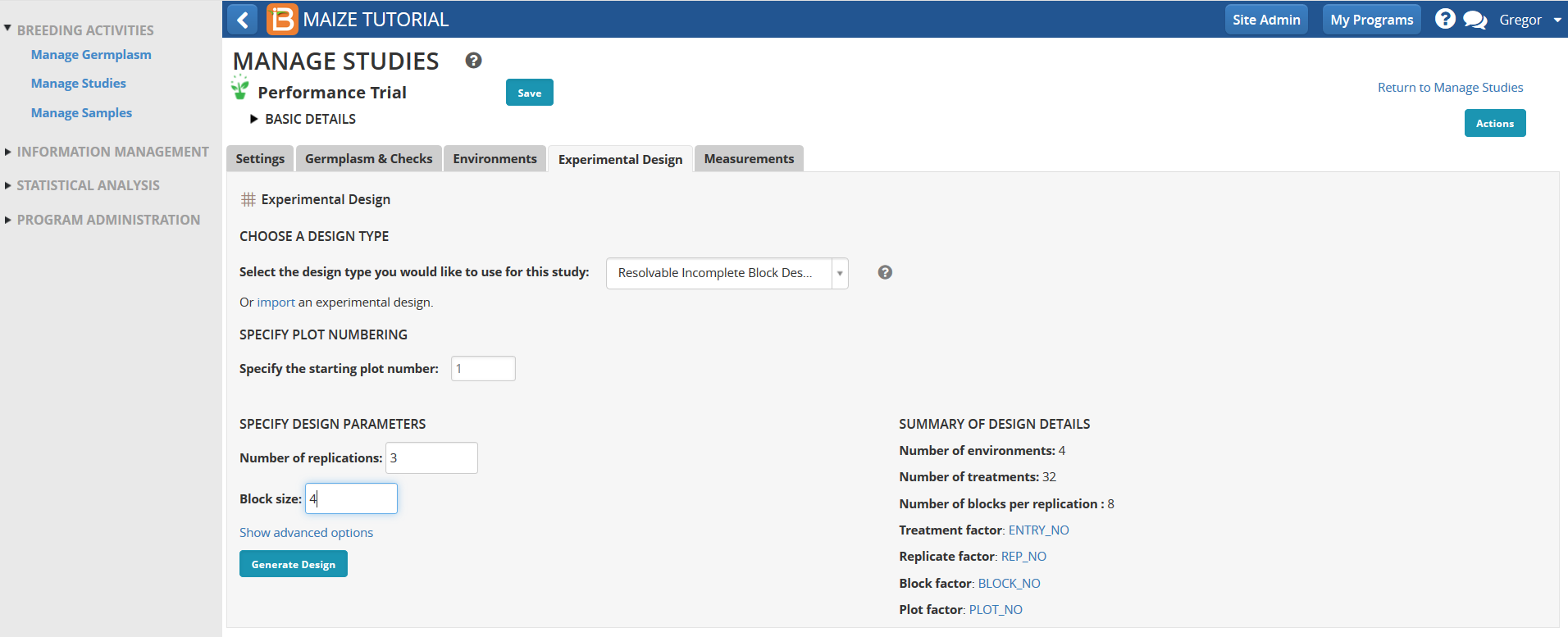
Import Trial Design
- Every study design generated by the BMS will have a different randomization. To use the tutorial example data, upload an experimental design file to match trial design to preformatted data import files.
Experimental Design File: Trial Design (.csv) - Export this Trial Design file and save it in your computer.
- Select import design from Design and planning options in the Actions Menu and browse to the saved Trial Design import file. Select Continue.
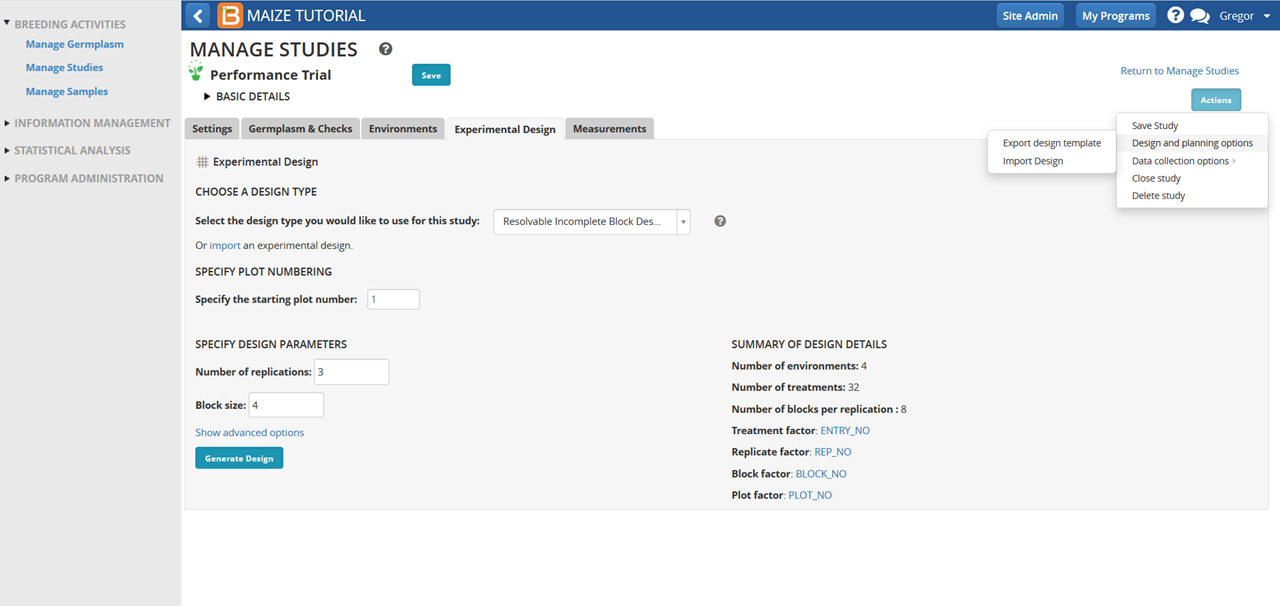
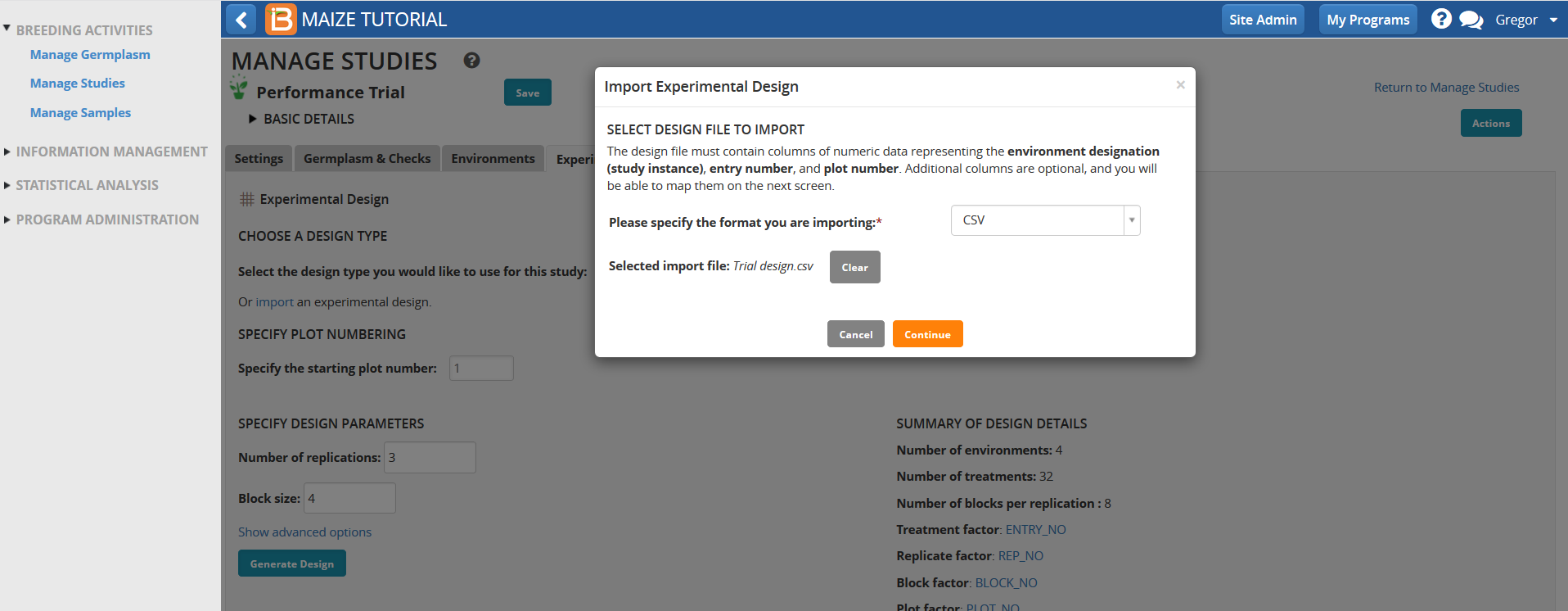
- Disregard the warning message. This experimental design is verified and compatible with BMS analysis tools. There are no unmapped variables. Select Next.
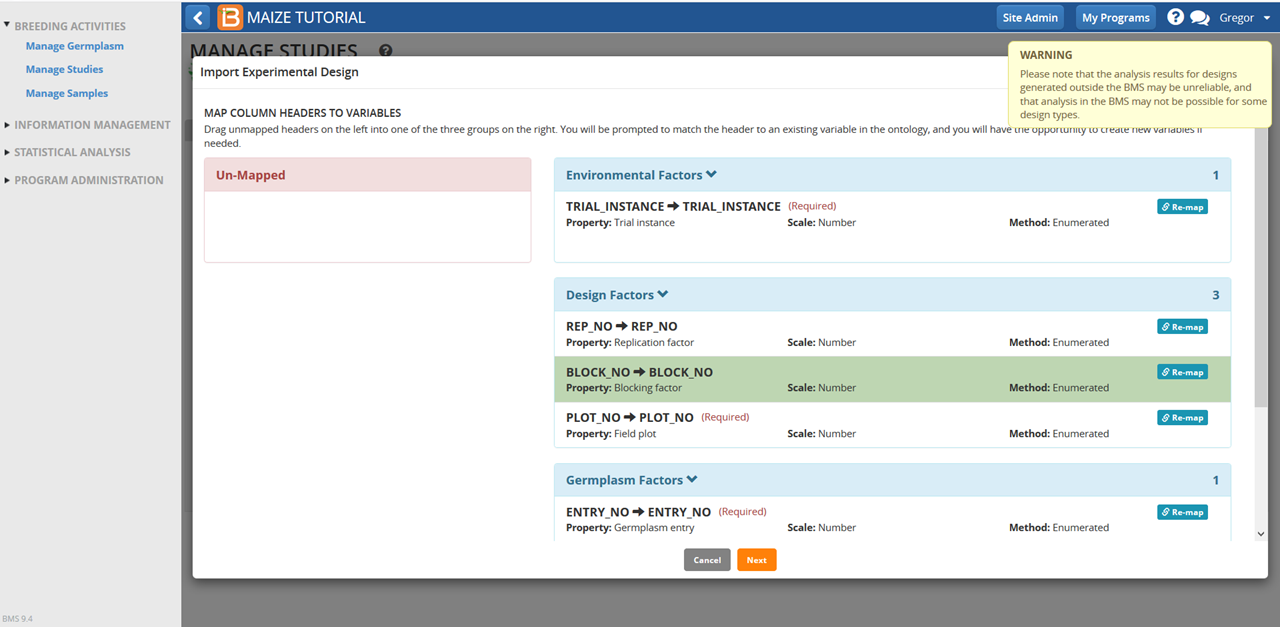
- If you have previously generated a study design, select Yes to overwrite the previous design.
- Review design details and select Finish at the bottom of the pop-up window.
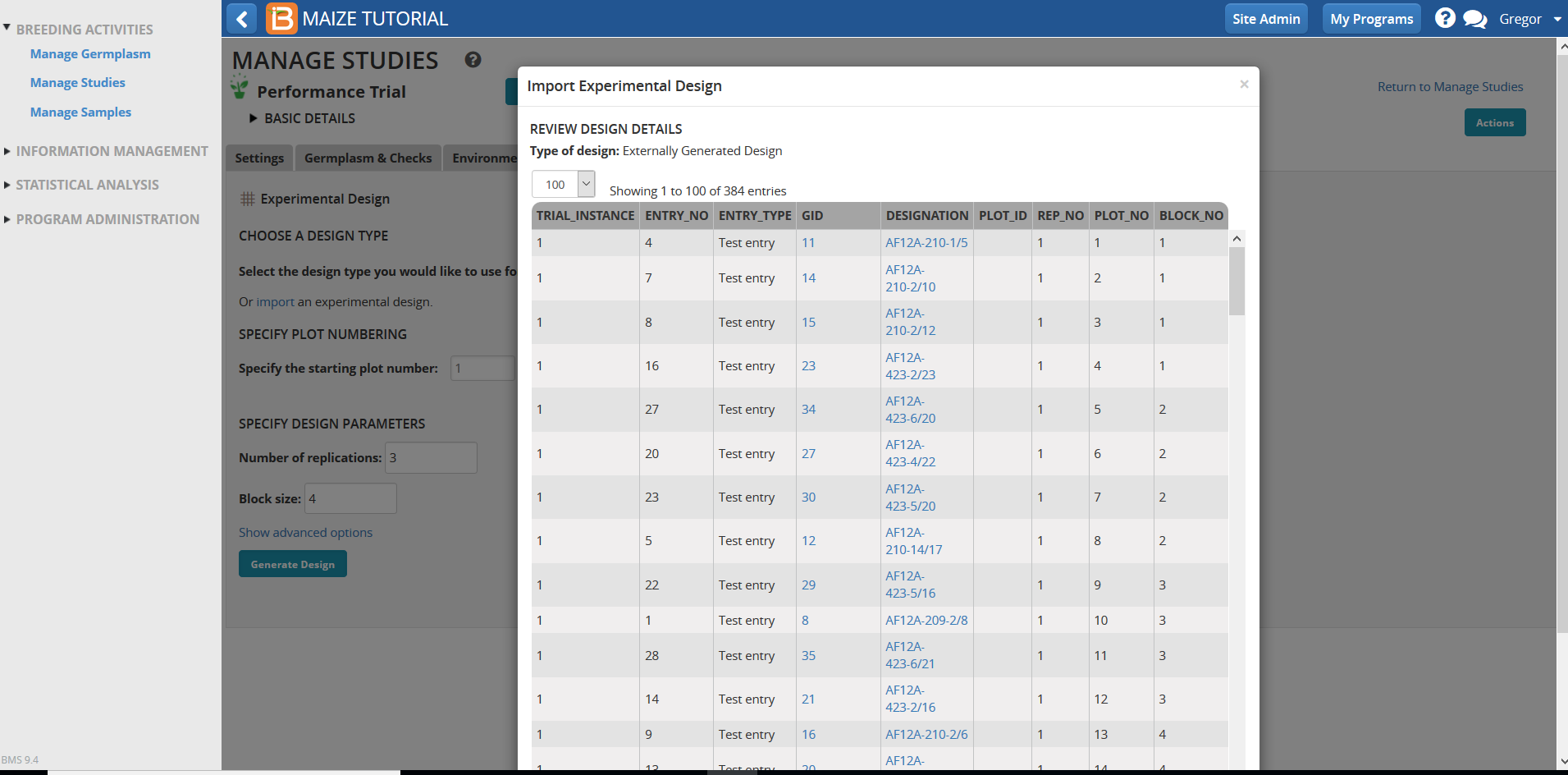
- Save and select the measurements tab.
Measurements
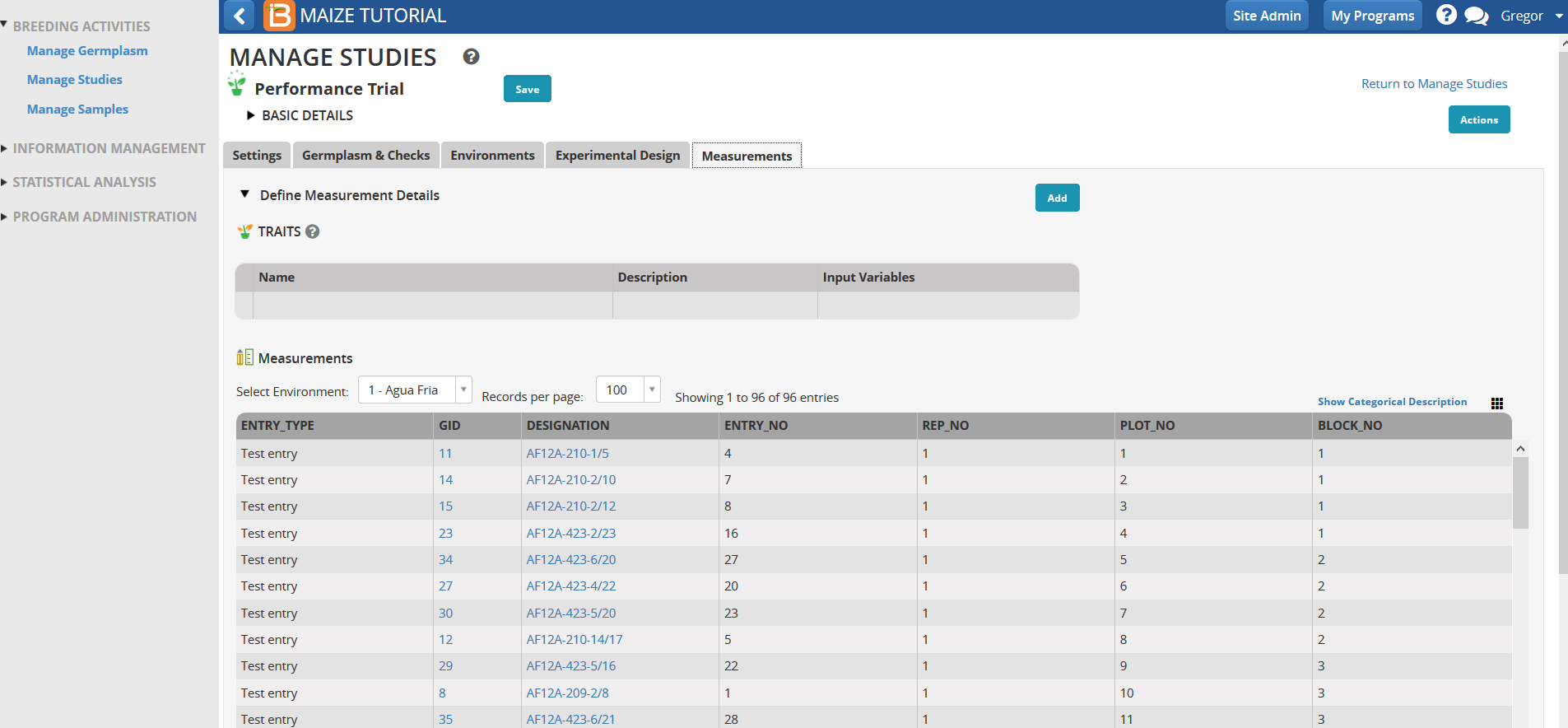
Once the trial design is saved, the measurements table is populated with independent variables and the dependent trait variables can be specified and added to the table.
This tutorial will collect and analyze data for six different traits.
- Add phenotypic traits by selecting Add associated with Traits.
- Search for the six traits by typing details of the individual traits in the selection bar. For example typing the first few letters of ‘Yield’ reveals all of the related measurements associated with yield.
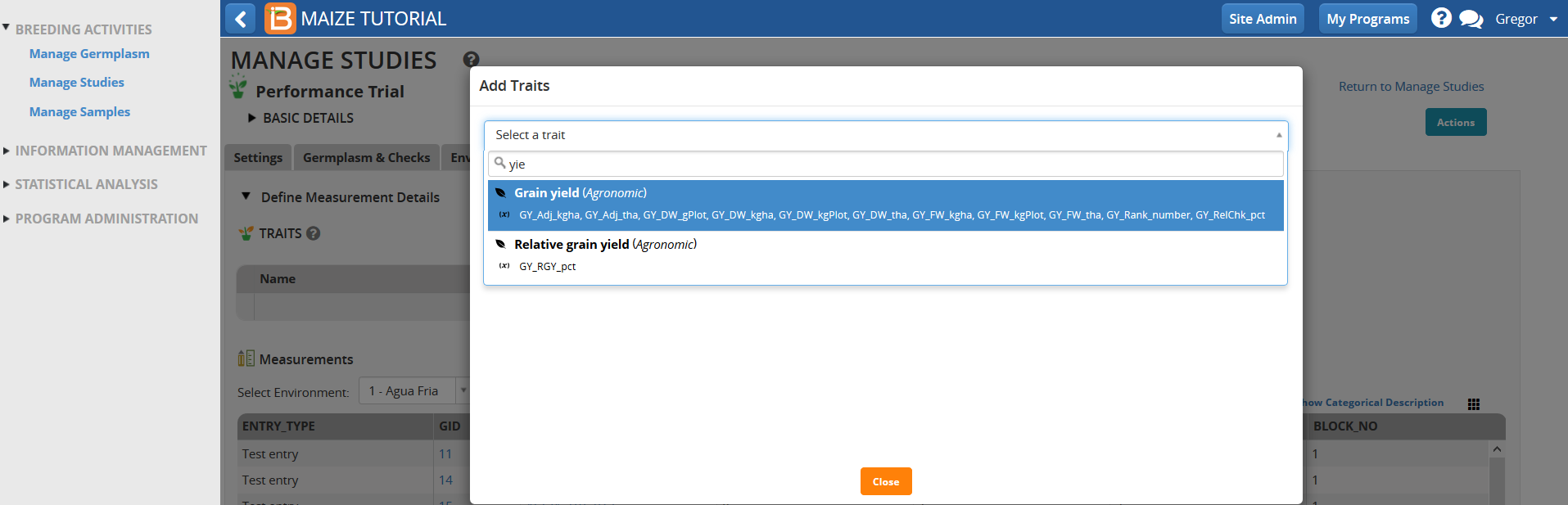
- Add the trait, GY_FW_kgPlot, (grain yield by fresh weight in Kg/plot) to the trial. Repeat by adding other traits indicated below.
- Grain Yield Fresh Weight (GY_FW_kgPlot,): Grain yield fresh in kg/plot
- Grain Moisture (GMoi_NIRS_pct): Grain moisture %
- Plant Height (PH_M_cm): Plant height to the insertion of the first tassel branch
- Ear Height (EH_M_cm): Ear height - Measurement in Cm
- Grain Yield Dry Weight (GY_DW_gPlot): Grain yield dry in g/plot
- Anthesis Date (Ant_DT_day): number of days from sowing to when 50% of the plants begin to shed pollen.
- Save the trial.
Note the added traits in the table.
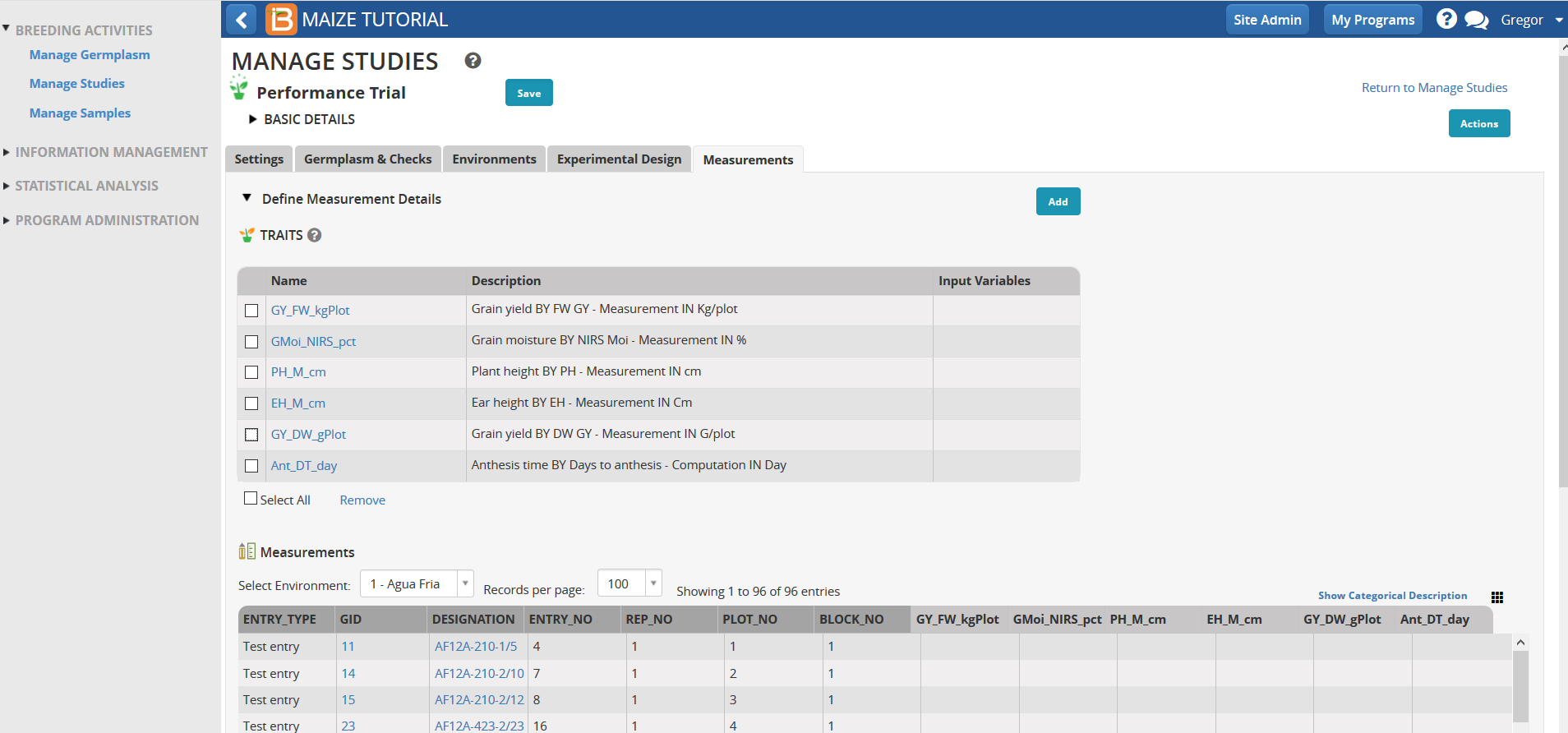
Create Planting Labels (Optional)
Trial design is complete when the measurements table is saved. Trial labels are available for seed packing and plot marking.
- Select Create Planting Labels under Design and Planning options in Actions menu.
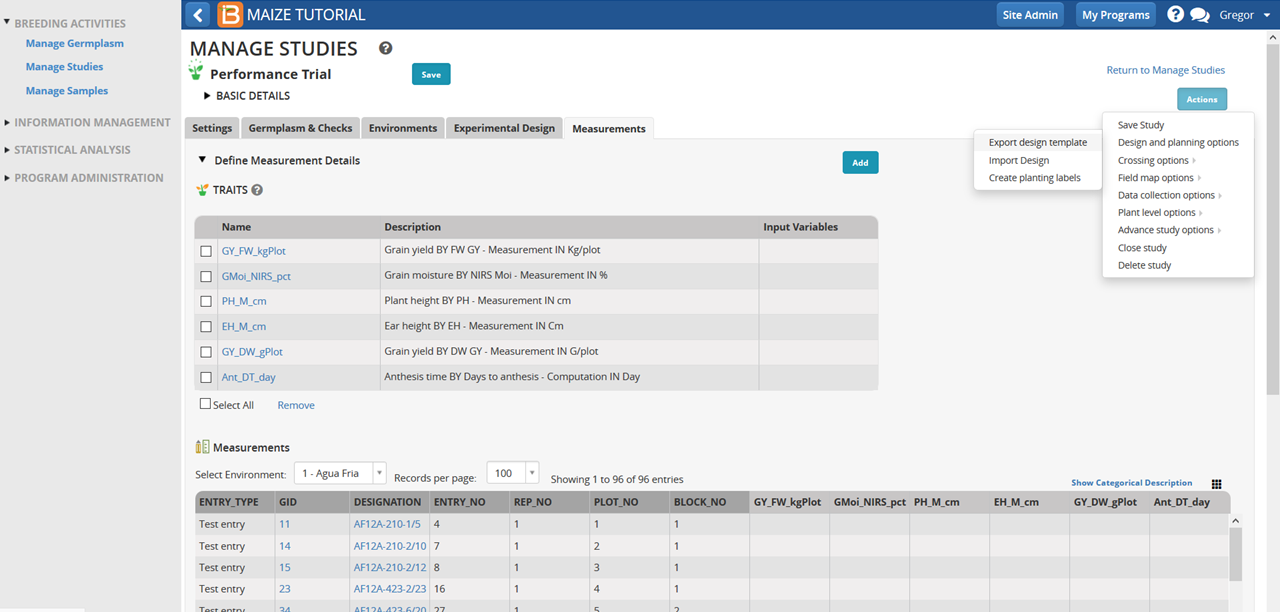
- Choose Excel as the output format. Excel format is a suitable input for a variety of label-making software options, including Bartender or Word (see video on using Word with BMS to create labels with barcodes).
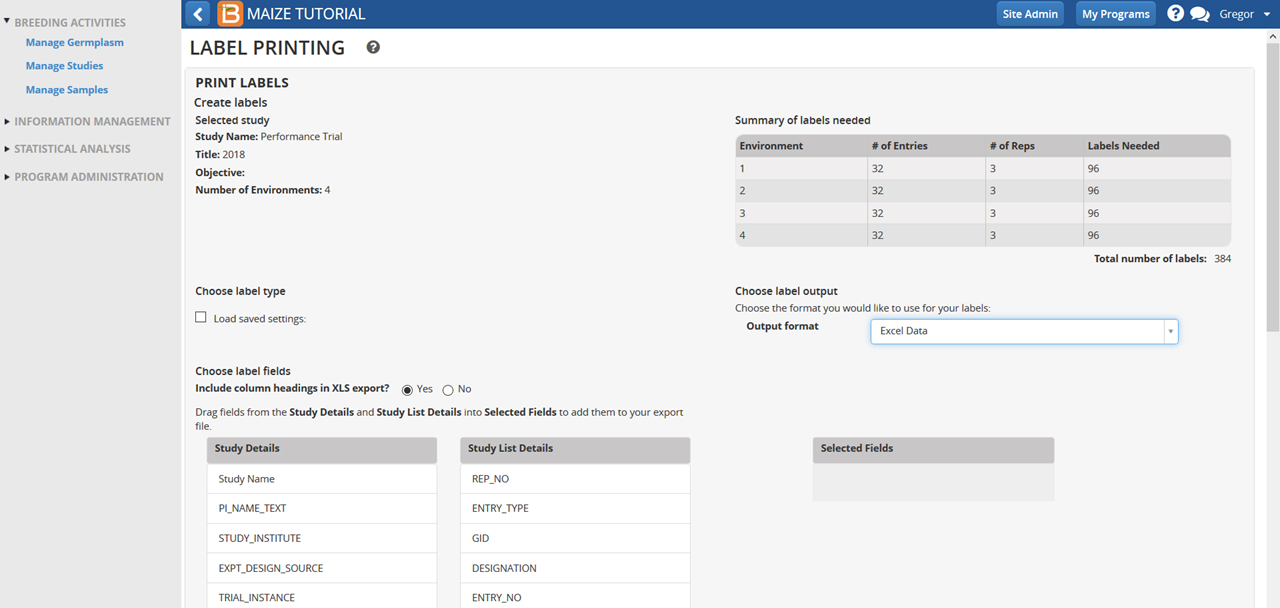
- Choose details to include on label text by dragging and dropping fields on Selected Fields.
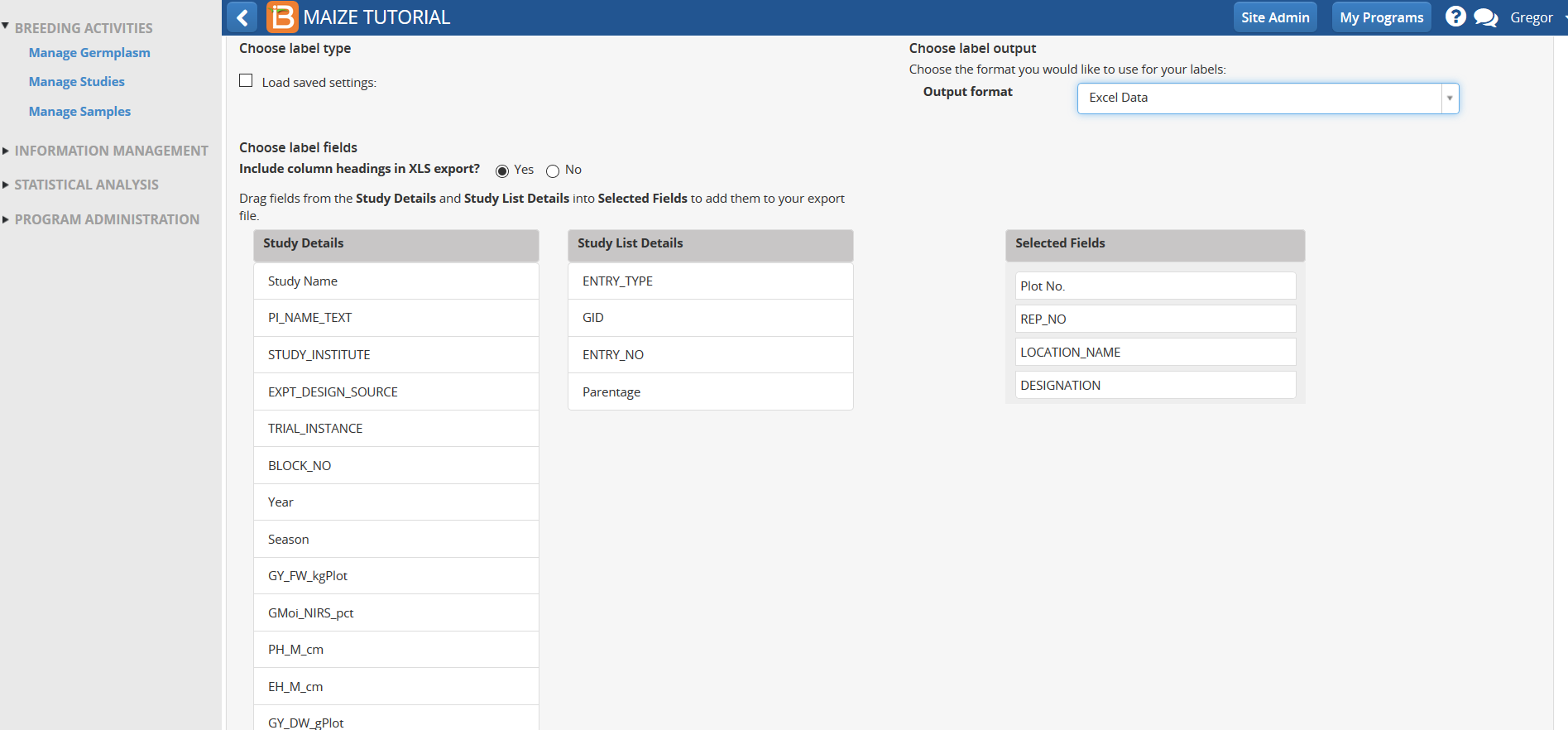
- Select Yes in the Barcode options
- Barcode your labels with unique plot id. Select Export label.
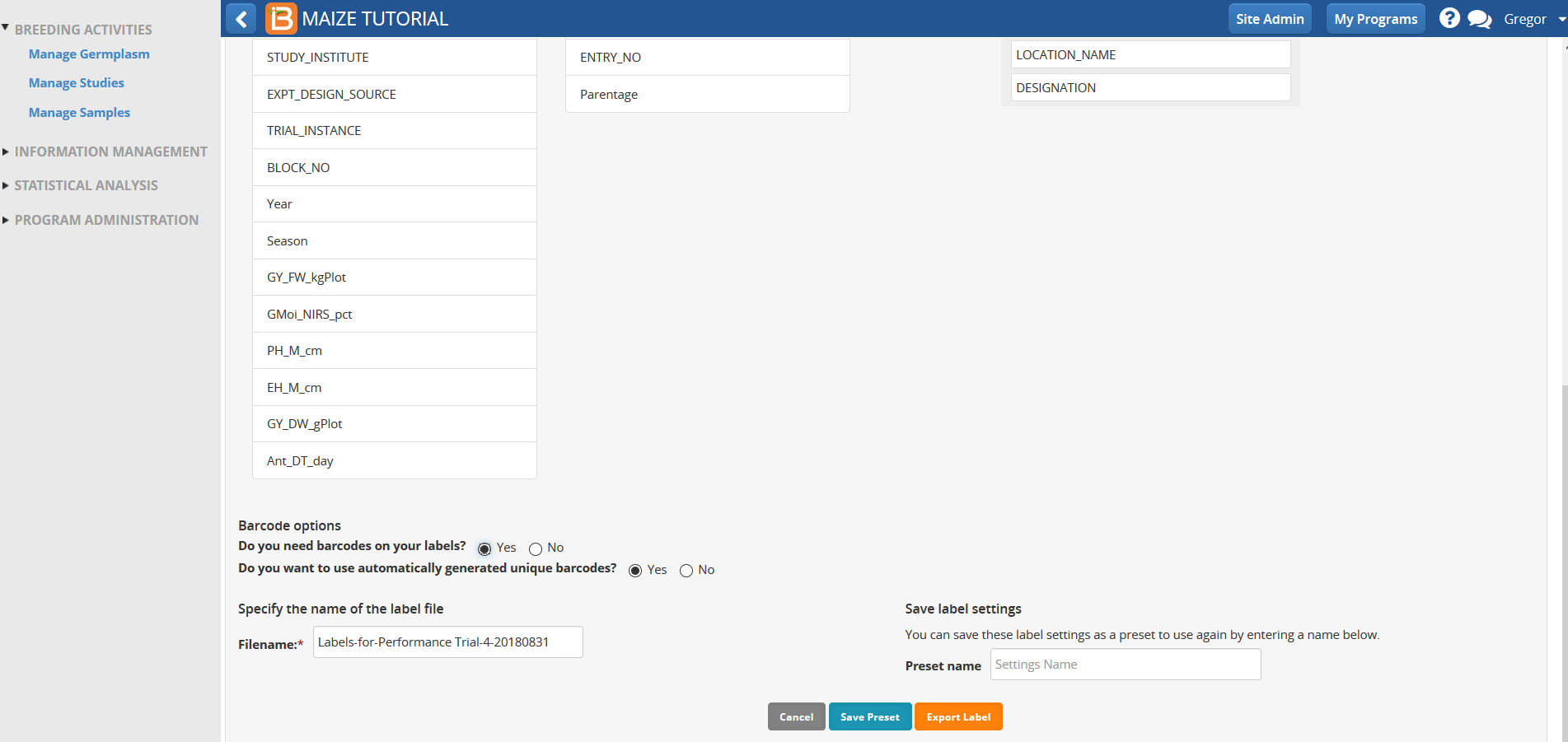
- Proceed without saving the label settings.
- Review the .xls file. The barcode is a unique PLOT_ID, which matches plot to seed inventory and phenotypic observations.
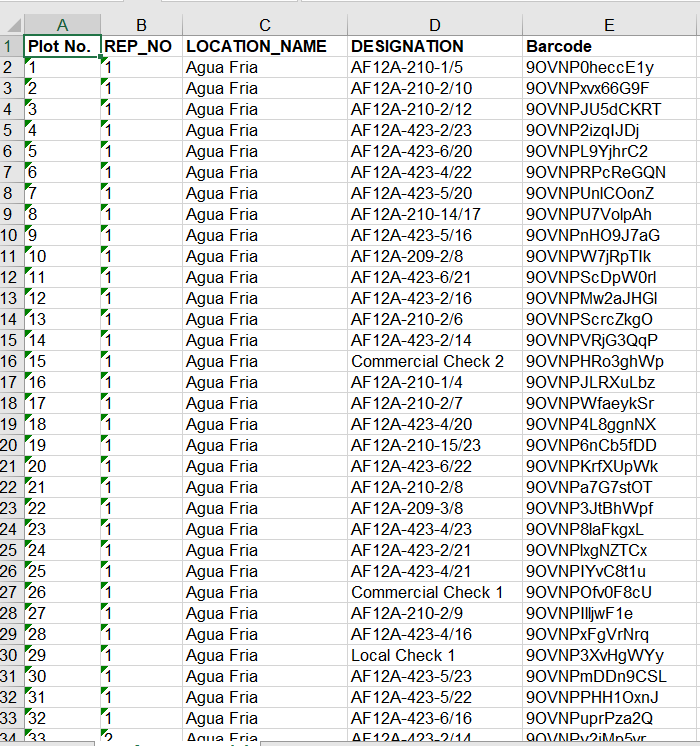
- Create labels with your label making software. The set of trial labels can be printed twice to label: the seed destined for the trial and (2) the trial plots in the field.

Reserve & Withdraw Trial Germplasm: Optional
If 40g of seed are needed per trial plot, and each trial germplasm is replicated 3 times over 4 environments - each trial germplasm entry requires a reservation and withdrawal of 480g of seed. Trial labels can be used to track and match packaged seed to appropriate plots.
- From Manage germplasm tool, browse and select the Trial Germplasm list. Close browse list popup window.
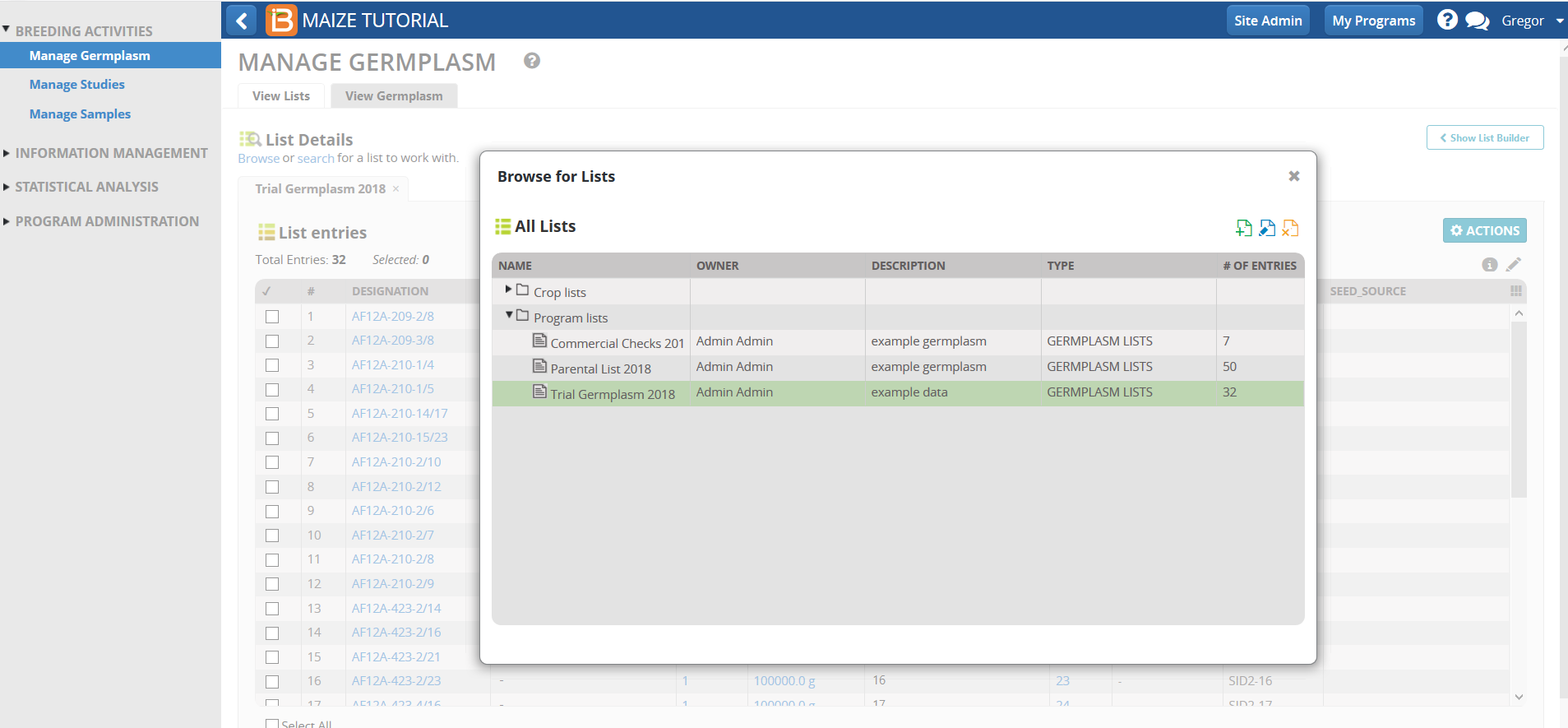
- Open the list in Inventory View.
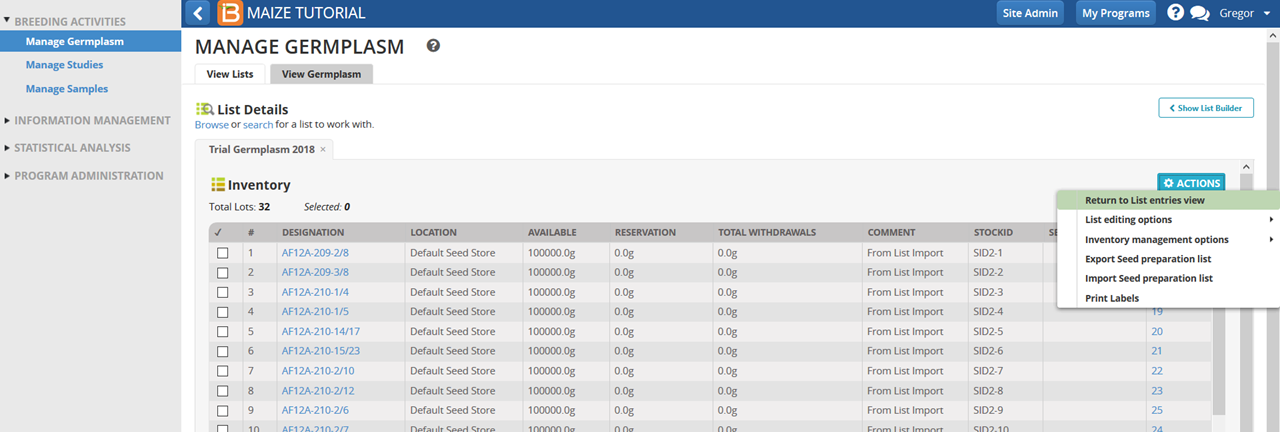
- Select all 32 entries and select Reserve Inventory.
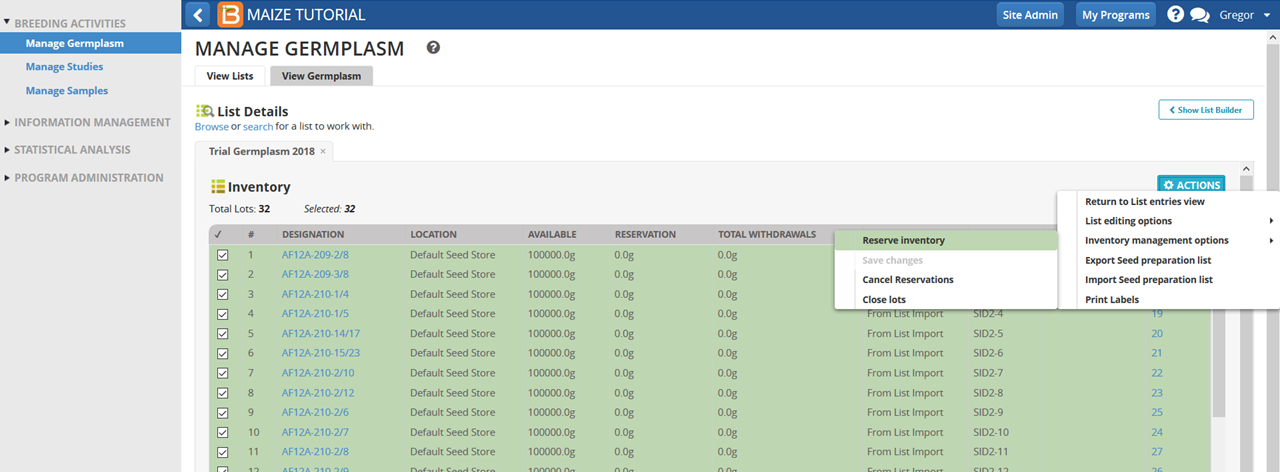
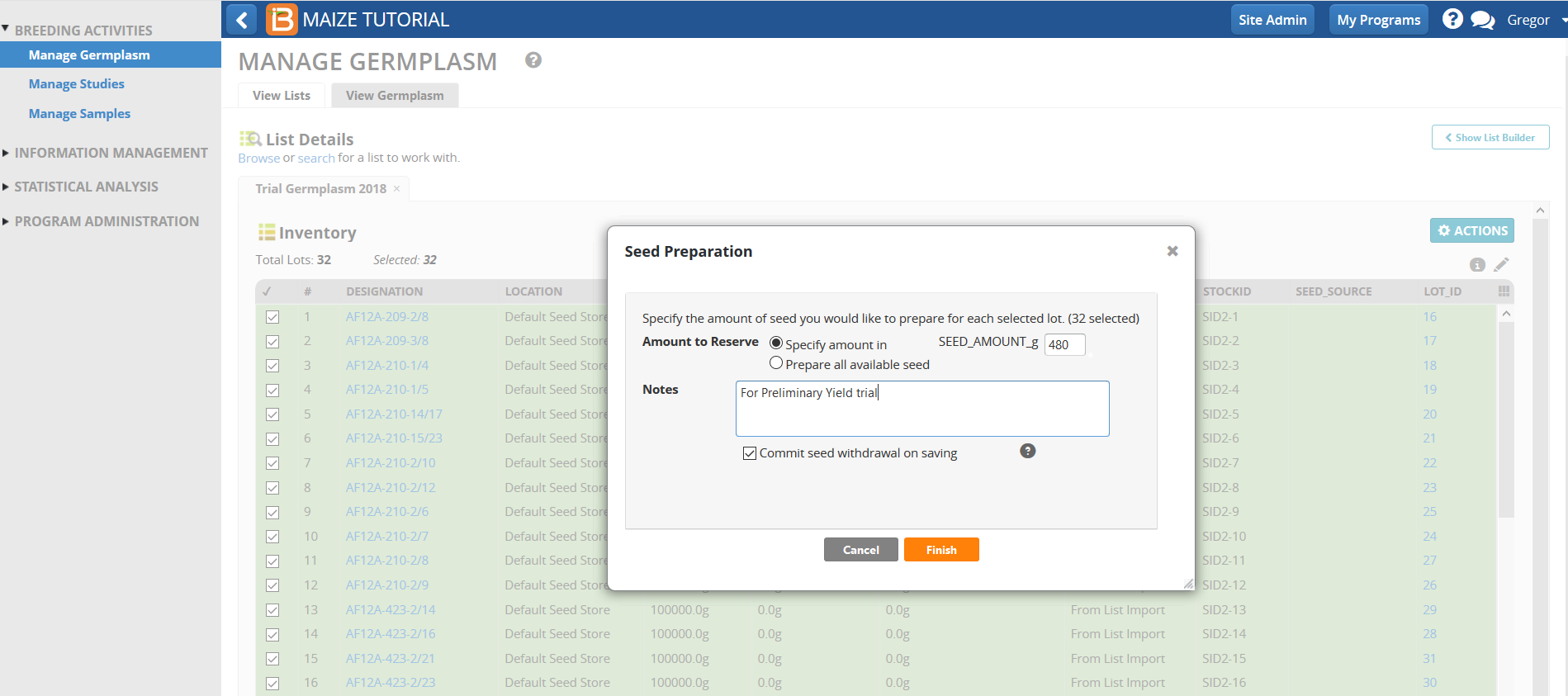
- Enter 480g for seed amount. Select Commit seed withdrawal on saving to complete the reservation/withdrawal in one step. Select Finish.
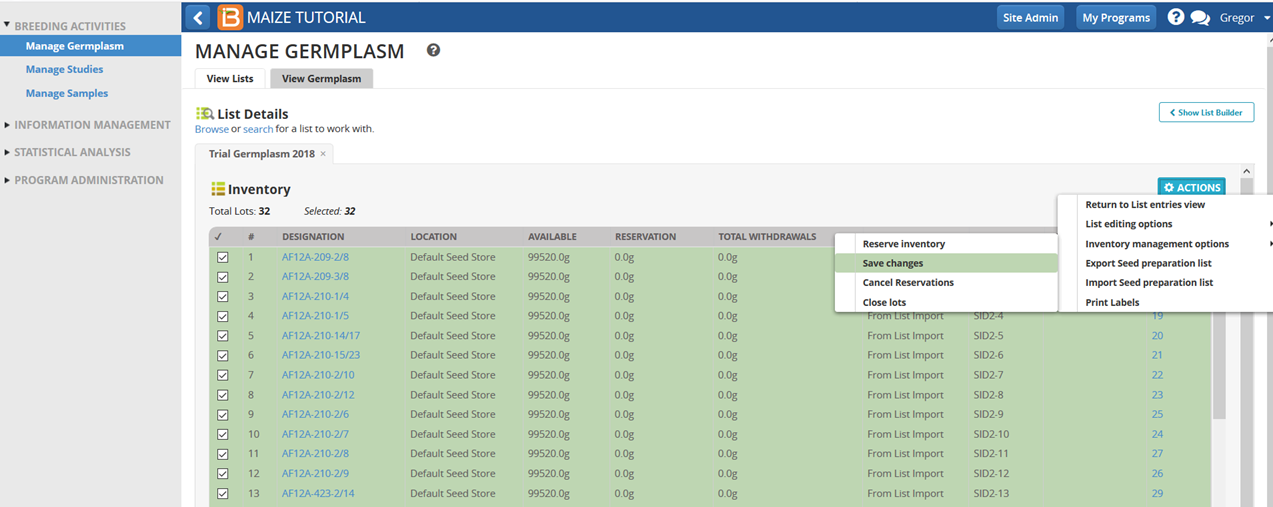
- Save changes.
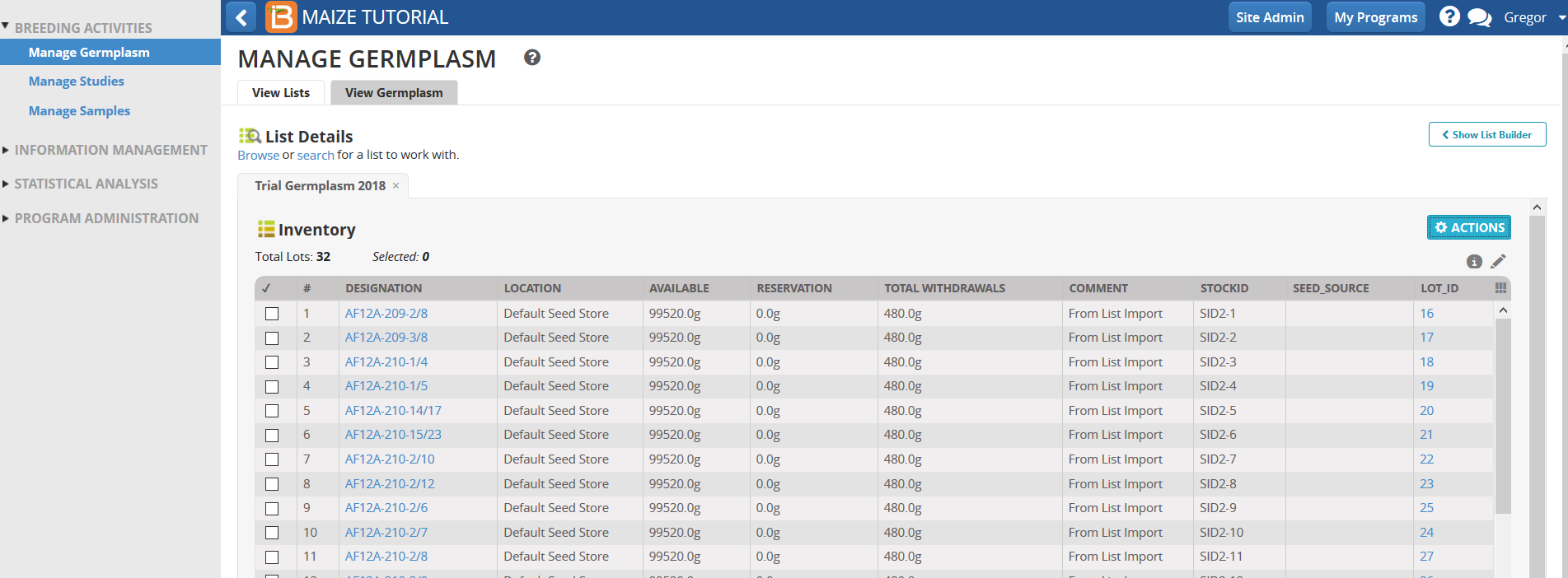
If you are working alone, TOTAL WITHDRAWALS will reflect your 480g withdrawal for each entry. If you are in a workshop with concurrent users, the total withdrawn by everyone will be reflected. When all the seed is used up or reserved, you will no longer be able to make reservations and withdrawals.
Related Materials
BMS Manual
Manage Trials
Install Breeding View
Maize Tutorials
Trial Data Collection
Single Site Analysis
Multi-Site Analysis
The Integrated Breeding Platform (IBP) is jointly funded by: the Bill and Melinda Gates Foundation, the European Commission, United Kingdom's Department for International Development, CGIAR, the Swiss Agency for Development and Cooperation, and the CGIAR Fund Council. Coordinated by the Generation Challenge Program the Integrated Breeding Platform represents a diverse group of partners; including CGIAR Centers, national agricultural research institutes, and universities.
Maize ?demonstration data was provided by Mike Olsen at CIMMYT International Maize and Wheat Improvement Center. These data have been adapted for training purposes. Any misrepresentation of the raw breeding data is the solely the responsibility of the IBP.

This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License?